Nanotheranostics 2026; 10:75-92. doi:10.7150/ntno.132646 This volume Cite
Research Paper
A Glucose-Responsive CeO₂@GOx Nanozyme Embedded in Chitosan/PVA Hydrogel for Accelerated Diabetic Wound Healing: from Molecular Simulations to In Vivo Validation
1. Nanotechnology Engineering, Faculty of Advanced Technology and Multidiscipline, Universitas Airlangga, Surabaya 60115, Indonesia.
2. Airlangga Functional Nanomaterials Research Group, Faculty of Advanced Technology and Multidiscipline, Universitas Airlangga, Surabaya 60115, Indonesia.
3. Department of Biological Pharmacy, Faculty of Pharmacy, Universitas Padjadjaran, Sumedang, Indonesia.
4. Department of Biology, Faculty of Mathematics and Natural Sciences, Universitas Indonesia, Indonesia.
5. Center of Excellence for Pharmaceutical Care Innovation, Faculty of Pharmacy, Universitas Padjadjaran, Sumedang, Indonesia.
6. Center for Sustainable Nanomaterials (CSNano), Ibnu Sina Institute for Scientific and Industrial Research (ISI- ISIR), Universiti Teknologi Malaysia, 81310 Johor, Malaysia.
7. Research Center for Computing, National Research and Innovation Agency, Jl. Raya Jakarta Bogor KM 46, Cibinong, Indonesia.
Received 2026-2-4; Accepted 2026-2-28; Published 2026-3-17
Abstract
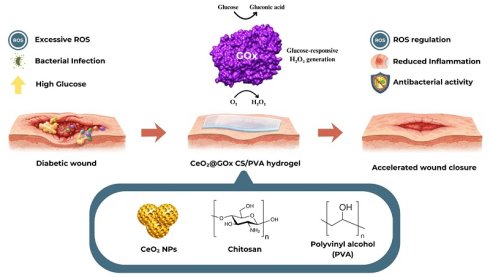
Diabetes prevalence in Indonesia reached 19.465 million patients in 2021 and is predicted to increase to 40.7 million cases by 2045, with 15-25% of diabetic patients at risk of diabetic wounds and 15-30% threatened with amputation. This study aims to determine the optimal concentration of cerium oxide nanoparticles functionalized with glucose oxidase (CeO2@GOx) nanozymes for the treatment of diabetic wounds. CeO2@GOx was incorporated into chitosan/poly(vinyl alcohol) (CS/PVA) hydrogel composites via a freeze-thawing method and systematically evaluated through physicochemical characterization, enzymatic assays, in silico analysis, antibacterial in vitro and in vivo wound healing studies. Molecular dynamics simulations (200 ns) were employed to investigate the dynamic stability of the flavin adenine dinucleotide cofactor within the GOx active site (FAD@GOx), particularly its structural persistence in the presence of Ce atom, CS, and PVA. Complementary density functional theory (DFT) calculations were independently performed on Ce₄O₈ clusters, chitosan, and poly(vinyl alcohol) (PVA) to examine their electronic properties and noncovalent interaction characteristics. UV-Vis analysis revealed increased absorbance at 298 nm with increasing CeO2@GOx concentrations, while functional group analysis confirmed the presence of hydroxyl, amine, carbonyl, and cerium-related features. The hydrogel exhibited a uniform particle size distribution (PDI = 0.1703) and a degradation rate of 90.334%. Among the tested formulations, the CeO2@GOx 4:1 (w/w) hydrogel exhibited the highest TMB oxidation rate and catalase-like activity and demonstrated optimal diabetic wound-healing performance in vivo within 7 days.
Keywords: Diabetes, Cerium nanozyme, Chitosan, hydrogel, wound healing
Introduction
Diabetes mellitus is defined as a condition where the body experiences metabolic abnormalities so that it is unable to produce insulin or insulin resistance [1-3]. Insulin is a hormone that balances blood glucose level. When the body lacks insulin, blood glucose levels will increase and interfere with the body's metabolism. Fasting blood glucose levels in people with diabetes can reach 125 mg/dL and above, with normal levels around 70-100 mg/dL and below [4]. Various health problems can arise due to diabetes, one of which is the risk of diabetic ulcer. Elevated blood glucose level can cause damage to tissues and blood vessels, potentially resulting in recurrent wounds. Consequently, the impaired circulation compromises the wound healing process, making it significantly more challenging for diabetic subjects to recover from such injuries [5,6].
In 2021, according to data from the International Diabetes Federation (IDF), Indonesia ranked seventh in the world for diabetes prevalence with 19.465 million people with diabetes. In addition, IDF also predicts that in 2045 diabetes will occupy the seventh position of the disease that causes death and the prevalence of diabetes is estimated to increase to reach 40.7 million people [7]. Diabetic wound complications are not uncommon in people with diabetes, 15-25% of diabetic patients experience diabetic wounds and with the increase in diabetes cases each year, diabetic wound cases will also increase [8]. About 15-30% of diabetic wound cases are threatened with amputation and various other life-threatening effects [9,10]. Diabetic wound management is challenging because wounds are more sensitive to infection, which is prevented by the administration of antimicrobial drugs either topically or by injection. However, the administration of antimicrobial drugs has not shown optimal results in protecting wounds from infection. Additionally, there is an urgency to develop a therapeutic agent that not only protect diabetic wounds from possible infection, but also reduce oxidative stress to accelerate wound healing.
Based on previous research, cerium oxide nanoparticles are able to mimic the concept of enzymatic activities (nanozymes), starting from superoxide dismutase (SOD), peroxidase, and catalase (CAT) enzymes that effectively break down excess reactive oxygen species (ROS) [11]. Glucose oxidase (GOx) can convert glucose and change it into gluconic acid and produce a by-product in the form of hydrogen peroxide (H2O2), so that glucose concentration is regulated [12,13]. This catalytic process is mediated by flavin adenine dinucleotide (FAD), which serves as the redox-active cofactor at the active site of GOx and plays a crucial role in electron transfer during glucose oxidation [14-16]. Previous studies have reported that glucose-triggered cascade reactions represent an emerging therapeutic strategy for wound healing, in which GOx-generated H₂O₂ can be further regulated or decomposed by nanozymes to maintain a balanced wound microenvironment [17]. Then the hydrogel composite is utilized as a topical application of the material on the wound. The use of chitosan as a material for making hydrogels is being widely researched, because the physicochemical properties of hydrogels can be designed to be responsive to certain external stimuli. In hydrogel fabrication, polyvinyl alcohol (PVA) is also used, which has a water absorption capacity 5-10 times its weight [18].
From the description of the benefits of each component above, this research utilizes cerium oxide nanoparticles functionalized with glucose oxidase (CeO2@GOx) in chitosan and PVA-based hydrogel composites (CS/PVA) as diabetic wound healing agents [19,20]. The characterization carried out includes the spectrum absorbance using Ultraviolet-Visible spectrophotometer (UV-Vis), functional group characterization using Fourier Transform Infrared Spectroscopy (FT-IR), Particle Size Analyzer (PSA) for the dispersity and stability test, hydrogel degradation test, enzymatic peroxidase and catalase tests, in silico analysis, antibacterial in vitro assay, and in vivo tests. This research will underpin an accessible yet effective treatment for diabetic wounds that supports Sustainable Development Goals (SDGs) point 3 of healthy living and well-being.
Materials and Methods
Materials
The research materials used include cerium (III) nitrate hexahydrate (Ce(NO3)3.6H2O) (Merck, Germany), citric acid (C6H10O8) (Merck), glucose oxide (GOx) from Aspergillus Niger (Chatson Jaya Lab, Indonesia), ethylene diamine tetraacetic acid (EDTA) (Merck), aquadest (H2O), ammonium hydroxide (NH4OH) (Merck, Germany), ethanol (C2H6O), (3-Aminopropyl)triethoxysilane (C9H23NO3Si) (Sigma Aldrich), phosphate buffer saline (PBS) pH 7,4 (Nitra Kimia), chitosan (CS) (Sigma Aldrich, USA), polyvinyl alcohol (PVA) (Sigma Aldrich, USA), acetic acid (CH3COOH) (Supelco, Germany), Tetraethyl orthosilicate (TEOS) (Sigma Aldrich), aquadest, hydrogen peroxide (H2O2), tetramethylbenzidine (TMB) (Merck), glutaraldehyde (C5H8O2) (Merck), citrate buffer, alcohol, ketamine (C13H16ClNO), and streptozotocin (STZ) (Chem Cruz, USA).
Methods of functionalization of CeO2@GOx CS/PVA
Density Functional Theory (DFT)
Density functional theory (DFT) calculations using ORCA version 6.0.1 were employed to investigate the electronic structure and noncovalent interactions features of representative molecular models. Geometry optimizations were performed using the hybrid PBE0 exchange-correlation functional together with the def2-TZVP basis set [21,22]. Long-range dispersion effects were accounted for by applying Grimme's D3 correction with Becke-Johnson damping (D3BJ) [22,23]. To improve computational efficiency, the resolution of identity approximation within the RIJCOSX scheme was adopted in conjunction with the def2/J auxiliary basis set [22]. Tight self-consistent field (TightSCF) convergence criteria were applied throughout all calculations [24,25]. CS and PVA were represented by short molecular segments to describe their local electronic environments. Based on previous theoretical studies reporting stable Ce-O coordination in Ce₄O₈ cluster [26]. In this research, Ce₄O₈ cluster was adopted to represent the local electronic environment of cerium oxide.
Molecular dynamics simulation
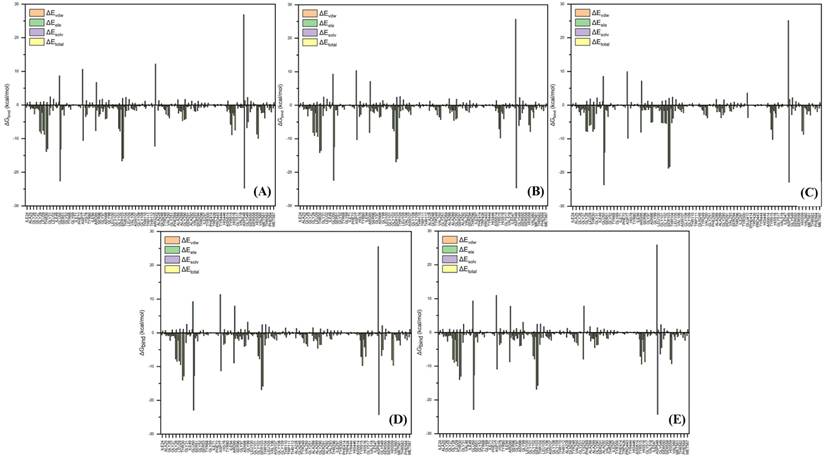
The 3D structure of glucose oxidase (GOx) was obtained from the Protein Data Bank (PDB ID: 1GAL). Protein modeling and reconstruction of missing residues were performed using the SWISS-MODEL (https://swissmodel.expasy.org/) (Fig. S1 and Fig. S2) [27]. The protein was identified using the PrankWeb (https://prankweb.cz/) to obtain active site of the protein (Table S1) [28]. Molecular docking was employed solely to generate initial protein-ligand conformations for subsequent molecular dynamics simulations, with the docking region defined around the FAD binding site. Docking calculations were performed using the AutoDock Vina package [29], while Ce atom, chitosan and polyviniyl alcohol docked were positioned within the predicted active site region. The docking grid box was defined with center coordinates (X: 47.9287 Å, Y: 15.4710 Å, Z: 58.6315 Å) and box dimensions (X: 14.479 Å, Y: 47.8644 Å, Z: 22.9982 Å). Docking poses were analyzed and visualized using UCSF Chimera software [30]. Molecular dynamics simulations were performed using the GROMACS 2023.02 package based on the CHARMM Generalized Force Field (CGENFF) [31]. The FAD@GOx complex obtained from the docking stage was used as the initial structure. The system was described using addition Ce atom parameter for CHARMM-All36 forcefield [32], with appropriate parameters assigned to the protein and the FAD cofactor. Energy minimization was conducted to remove steric clashes, followed by equilibration under constant volume (NVT) and constant pressure (NPT) conditions at 310 K and 1 atm for a a total of 4 ns. A production MD simulation was then carried out for 200 ns under periodic boundary conditions. Binding free energy calculations were carried out using the Molecular Mechanics/Generalized Born Surface Area (MM-GBSA) approach implemented in the cpptraj and MMPBSA.py [33]. The binding free energy (∆Gbind) was evaluated by combining the gas-phase interaction energy (∆Ggas) and the solvation free energy contributions (∆Gsolv). As described in Eqn. (1), the (∆Ggas) represents the interaction energy between the ligand and the receptor in the absence of solvent, obtained from the difference between the complex and the separated components. The (∆Gsolv) was calculated as shown in Eqn. (2). Furthermore, the solvation contribution was decomposed into electrostatic (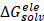 ) and nonpolar components (
) and nonpolar components (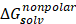 ), as defined in Eqn. (3).
), as defined in Eqn. (3).
(1)
(2)
(3)
Synthesis of CeO2 NPs
The synthesis of CeO2 NPs was performed by the precipitation method at 27°C. First, 10 mL of 1% EDTA solution, 10 mL solution of 1% C6H10O8 was added into 10 mL of Ce(NO3)3.6H2O 0.8 M. Then, add 50 mL of 0.8 M NH4OH dropwise and stirred at 450 rpm. After 24 hours of mixing, the samples were transferred into falcon tubes and left at room temperature for 2 days. The sample was centrifuged at 4,000 rpm for 20 minutes and the precipitate was collected. Then, the precipitate was washed using distilled water twice and ethanol once, then dried for 15 hours at 60°C. In the final stage, the sample was crushed to a fine yellow powder of CeO2 NPs.
Synthesis and immobilization of CeO2@GOx
The formation stage of CeO2@GOx begins with CeO2 NPs re-dispersed in 50 mL ethanol via sonication for 15 minutes. Then, 200 μL of 3-(aminopropyl)triethoxysilane was added, stirred at room temperature overnight. The precipitate was washed with 3x4 mL of ethanol and dried in air at room temperature. The amino-functionalized CeO2 NPs were then washed with 4x4 mL PBS and re-suspended in 5 mL PBS solution. On the other hand, GOx was dissolved into 2 mL of PBS solution pH 7.4 and incorporated into the CeO2 NPs solution. The reaction mixture was gently stirred overnight. The CeO2@GOx solution was washed with 4 times 4 PBS solution pH 7.4. The solution was resuspended in 5 mL of PBS solution pH 7.4.
Synthesis of CeO2@GOx and incorporation into CS/PVA hydrogel composites
CS/PVA hydrogel was synthesized by first dissolving CS in 1% (w/v) acetic acid at room temperature for 24 h under constant stirring. In parallel, PVA was dissolved in distilled water (10% w/v) at 90-99°C until a homogeneous solution was obtained. Subsequently, the PVA solution was added to the CS solution, followed by the addition of 2 mL of TEOS. The mixture was stirred at 70°C for 3 h until homogeneous. After cooling to room temperature, the solution was ultrasonicated for 5 min without temperature control. Separately, the CeO2@GOx solution was suspended in PBS and sonicated for 10 min. The CeO2@GOx suspension was then added dropwise into the CS/PVA solution under continuous stirring for 2 h. The resulting mixture was cast into a petri dish and allowed to rest at room temperature for 30 min. The petri dish was covered with aluminium foil and left undisturbed for 24 h. The formed hydrogel was then transferred into a microtube and subjected to a freeze-thaw process at -20°C until completely frozen. After freezing, the hydrogel was allowed to thaw at room temperature, and its structure was subsequently observed [34,35].
Characterization of CeO2@GOx CS/PVA
Spectrophotometric evaluation using UV-Vis
Ultraviolet-Visible (UV-Vis) is a spectroscopic technique commonly used for the measurement of light absorbance at various wavelengths that are still in the ultraviolet (UV) range (190-400 nm) and visible (Vis) range (400-800 nm) derived from the electromagnetic spectrum [36]. The UV-Vis test was used to determine the responsiveness of the nanozyme sample to the absorbance of CeO2 NPs and GOx respectively in the CeO2@GOx sample variation using the GENESYS 180 UV-Vis Spectrophotometer, where all samples were in liquid form for UV-Vis analysis. In addition, this test is also used to determine the absorbance intensity when there is the addition of CeO2 NPs in CeO2@GOx [37,38].
Functional group characterization using FT-IR
Functional group characterization was carried out using fourier transform infrared (FT-IR) Spectroscopy is a tool to analyze a functional group in a chemical compound [39]. The FT-IR test used The PerkinElmer Spectrum™ 3 FT-IR spectrometer with a wave number range from 4000-400 cm-1. The samples characterized were in liquid form, including CeO2@GOx, CS/PVA, and all variations of the variation of hydrogel samples such as CeO2@GOx CS/PVA 1:1, CeO2@GOx CS/PVA 2:1, and CeO2@GOx CS/PVA 4:1. Prior to measurement, the FT-IR detection area was thoroughly cleaned with ethanol to eliminate residual impurities and minimize background interference during spectra acquisition.
Particle size diameter test
The particle size diameter test employs dynamic light scattering (DLS) to assess the dispersion of a system [40,41]. Particle size analysis (PSA) typically measures the Brownian motion of individual particles in a liquid to analyze their size. The PSA assay was utilized to evaluate the dispersity of CeO2 NPs and CeO2@GOx, and to measure their particle size diameter using a BIOBASE BK-802N device, which measures in the range of 1-1000 nm.
Degradation test
The degradation test with PBS was performed to simulate the hydrogel's behaviour in the body when used as a diabetic wound dressing [34]. The degradation test was carried out by immersing all samples in PBS solution pH 7.4 and left in a room around 25°C for 192 hours or 8 days. To evaluate the physical resistance of the samples in degradation, the samples were weighed every day to measure the percent mass ratio of the initial mass of the samples using PBS. Degradation was carried out under the closest conditions for the interaction between hydrogel and skin in a biological environment [42], so PBS solution was chosen to facilitate this [34]. In the calculation of degradation, the following formula of Eqn. (4) was used:
(4)
with W as the mass per day and Wi as the initial mass of the sample.
Enzymatic test of CeO2@GOx
The peroxidase and catalase enzymatic assays were performed according to the research references with some special adjustments and modifications [37,43]. Catalase is one of the working nanozymes used to degrade H2O2, an oxidizing agent harmful to the diabetic wound environment [35]. Nanocerium shows catalase-like behavior, especially when the Ce+4/Ce+3 ratio is elevated, as evidenced by methods such as the Amplex Red assay and UV-Vis spectrophotometry, where H2O2 absorption is tracked at 240 nm. Peroxidases, enzymes typically characterized by the presence of a heme cofactor in their active sites, are prevalent in living organisms; glutathione peroxidase is one notable example [20]. Peroxidase and catalase mimetic activities utilize hydrogen peroxide as a substrate and break it down into harmless water and oxygen molecules in catalase activity and/or generate hydroxyl and perhydroxyl radical cascades in peroxidase activity at acidic pH values [43]. In enzymatic peroxidase reactions, substrates such as the colorless 3,3',5,5'-tetramethylbenzidine (TMB) undergo oxidation, resulting in the formation of a colored product in the presence of peroxidase [20].
Antimicrobial assay
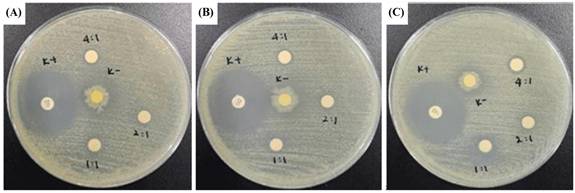
An antimicrobial assay was done to analyze the CeO2@GOx sample capabilities further to prevent possible wound bacterial infections. The antimicrobial assay was achieved through an inhibition zone test named disc diffusion assay (DDA). Gram-negative bacteria Escherichia coli ATCC 25922 (E. coli) and Gram-negative bacteria Staphylococcus aureus ATCC 25923 (S. aureus) were used in this test as the bacterial representative. Escherichia coli and Staphylococcus aureus were grown in NB (nutrient broth) media and incubated for 24 hours at 37℃. A 100 µl sample of 24-hour-old microorganisms, which had reached an OD = 0.1 in physiological water, was taken and spread on a Petri dish that had been pre-filled with MHA (Mueller Hinton Agar) media. MHA media was used to test antimicrobial samples against the growth of E. coli and S. aureus. The samples used in this test were in liquid to semi-gel form with the codes CeO2@GOx 1:1, 2:1, and 4:1 w/w. The samples were taken in a volume of 30 µl and dropped onto a sterile blank disc. The positive control used gentamicin as a commercial antimicrobial agent, and the negative control used aquadest. Then, the Petri dishes were incubated at 37℃ for 24 hours. The diameter of the inhibition zone formed around the disc containing the sample was measured using calipers. The blank disc used had a size of 5 mm [44,45].
In vivo study
In vivo tests were performed on male BALB/c strain mice (Mus musculus), 4 weeks of age, and body weight 20-30 grams, divided into four different treatment groups, namely the Normal Group (KN) without being given Streptozotocin (STZ), the Negative Group (K-) given STZ, the Positive Group (K+) given STZ and given conventional diabetic wound medication in the form of Betadine ointment, and the Treatment Group (KP) given a hydrogel composite with the best concentration based on characterization and a series of previous tests. STZ was administered at a dose of 30 mg/kg once a day, for five consecutive days based on multiple low-dose diabetes index. After induction, only the K-, K+, and KP groups were confirmed to have developed diabetic conditions and proceeded to treatment, while the KN group was injected with a citrate buffer to ensure all mice were subjected to equivalent stress conditions. After that, visual observation of wound healing was carried out on mice for 7 days [1]. The number of mice for 4 groups was adjusted according to Federer's formula, resulting in a minimum of 6 mice in 1 group and a total of 24 mice. The procedure of this study was approved by the Research Ethics Commission, Faculty of Veterinary Medicine, Airlangga University (No. 3.KEH.105.07.2024).
Results and Discussion
Density functional theory (DFT) study
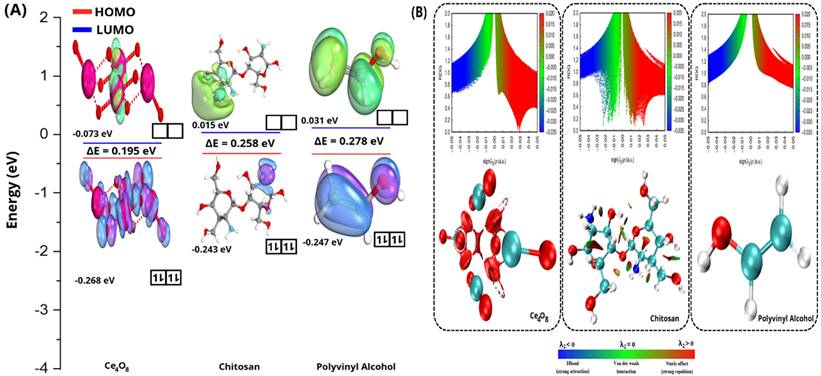
The electronic properties of Ce₄O₈ cluster, CS, and PVA were analyzed through HOMO-LUMO and Reduced Density Gradient (RDG) analyses (Figure 1A). The HOMO-LUMO distributions illustrate the electron-donating and electron-accepting regions of each system, while the energy gap (ΔE) reflects their electronic stability and reactivity [46,47]. Ce4O8 exhibits the smallest energy gap, indicating higher electronic reactivity compared to chitosan and PVA. In contrast, chitosan and PVA show relatively larger ΔE values, suggesting more stable electronic structures. In addition, the energies of the HOMO and LUMO were employed to derive global reactivity descriptors, including chemical hardness (η), softness (S), chemical potential (μ), electronegativity (χ), electrophilicity index (ω), and the maximum amount of electronic charge transfer (ΔNmax). These descriptors provide quantitative insight into the intrinsic electronic behavior of each system and presented in Table 1. The descriptors were calculated using Eqn. (5-10) based on established theoretical formulations reported in previous studies [48-50].
Electronic properties; (A) HOMO-LUMO with their energy levels and energy gaps (ΔE), and (B) RDG analysis.
Global reactivity descriptors of Ce₄O₈, CS, and PVA
| Descriptors | Ce4O8 | CS | PVA |
|---|---|---|---|
| Electronegativity (χ) (eV) | 0.171 | 0.114 | 0.108 |
| Chemical potential (μ) (eV) | -0.171 | -0.114 | -0.108 |
| Chemical Hardness (η) (eV) | 0.098 | 0.129 | 0.139 |
| Softness (S) (eV-1) | 5.102 | 3.876 | 3.597 |
| Electrophilicity index (ω) (eV) | 0.149 | 0.050 | 0.042 |
| ΔNmax | 1.745 | 0.884 | 0.777 |
(5)
(6)
(7)
(8)
(9)
(10)
The reduced density gradient (RDG) analysis was further employed to visualize and characterize the noncovalent interactions present in the Ce₄O₈ cluster, CS, and PVA (Figure 1B). The RDG plots, represented as scatter graphs of RDG versus sign(λ₂)ρ, enable the identification of different types of interactions. Peaks located at negative values of sign(λ₂)ρ correspond to attractive interactions such as hydrogen bonding, while regions around zero indicate van der Waals interactions, and positive values are associated with steric repulsion. Gradient isosurfaces offer an intuitive visualization of weak interactions as spatially extended regions, rather than as isolated pairwise atomic contacts [51]. When the reduced density gradient is mapped onto the molecular geometry, continuous surfaces appear, allowing the identification of green regions associated with attractive interactions and red regions corresponding to steric effects [51-53].
Molecular dynamics simulation study
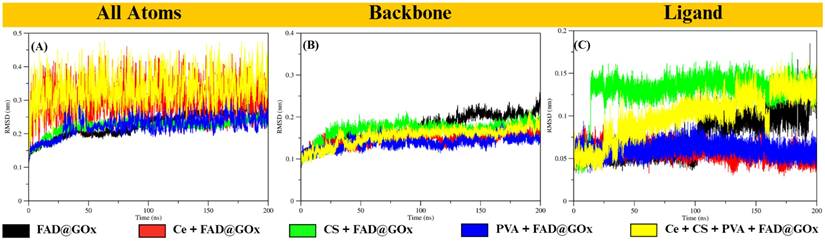
To evaluate the structural stability of GOx and the behavior of the FAD cofactor under different simulation conditions, molecular dynamics simulations were performed and analyzed using root mean square deviation (RMSD) analysis (Figure 2). The RMSD behavior observed in this study is consistent with previously reported molecular dynamics simulations of the GOx-FAD complex [33]. In earlier studies, GOx systems exhibited an initial increase in RMSD followed by stable fluctuations, which was attributed to conformational adjustment during equilibration while preserving the overall protein fold. Similarly, in our simulations, the all-atom RMSD in the Ce + FAD@GOx and Ce + CS + PVA + FAD@GOx systems increases from approximately 0.2 to 0.4 nm before reaching a stable plateau, indicating structural relaxation without significant unfolding. Notably, the RMSD values obtained in this work remain within a comparable or lower range than those reported for GOx-FAD and GOx-FAD-glucose complexes, suggesting that the incorporation of Ce atom, CS, and PVA maintains the global structural stability of the GOx structure. However, analysis of the Ce atom trajectory reveals that, over the 200 ns simulation, Ce tends to migrate away from the GOx surface, particularly in the presence of the CS-PVA matrix (Figure S3). This behavior indicates that Ce does not form persistent direct interactions with the protein, and its influence on GOx is likely indirect, mediated through the surrounding environment rather than strong binding to the active site. Overall, the preserved RMSD profiles confirm that the global fold of GOx remains stable throughout the simulation, while local flexibility-especially in the FAD-binding region is retained. The average RMSD values for all systems are summarized in Table S2.
RMSD analysis of FAD@GOx systems under different simulation conditions: (A) All atoms RMSD, (B) backbone, and (C) ligand RMSD with a specific focus on the FAD cofactor relative to the GOx structure.
To gain insight into the conformational stability of GOx and the dynamic behavior of its catalytic environment, the structural evolution of the FAD@GOx systems was examined during the production phase of the molecular dynamics simulations [54,55]. Particular attention was given to the preservation of the overall protein fold and the integrity of the FAD-binding pocket, as these features are critical for maintaining enzymatic function. Representative structural snapshots extracted from the later stages of the simulations (125-150 ns) are shown in Figure 3A, demonstrating the absence of large-scale unfolding or significant structural distortion among the different systems. These observations indicate that the catalytic microenvironment of GOx remains structurally intact despite variations in the surrounding simulation conditions. The residue of root mean square fluctuations (RMSF) profiles (Figure 3B) further reveal that the majority of GOx residues exhibit limited fluctuations, particularly in regions proximal to the FAD-binding pocket. The average RMSF values of the apo protein systems are summarized as follows: FAD@GOx (0.128 ± 0.003 nm), Ce + FAD@GOx (0.110 ± 0.002 nm), CS + FAD@GOx (0.116 ± 0.002 nm), PVA + FAD@GOx (0.105 ± 0.002 nm), and Ce + CS + PVA + FAD@GOx (0.118 ± 0.002 nm).
Structural and dynamic properties of FAD@GOx systems; (A) Representative structural snapshots at 125-150ns for simulations system, (B) RMSF profiles of FAD@GOx residues.
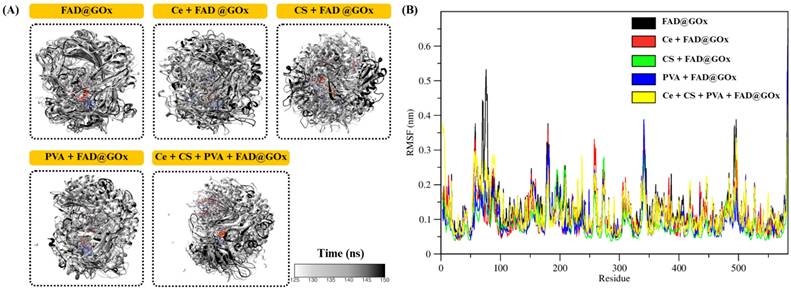
Importantly, key catalytic residues previously identified in the literature such as GLU412, HIS516, and HIS559, display relatively low RMSF values compared to surface-exposed and loop regions. This restrained flexibility suggests a stable catalytic microenvironment, which is essential for efficient electron and proton transfer during glucose oxidation [56]. This observation is consistent with previous studies that describe these residues as critical components of the GOx catalytic machinery, in which HIS516/520 and HIS559/563 function as potential proton acceptors, while GLU412 stabilizes the hydrogen-bond network near the FAD moiety [57,58]. Consistent with prior structural and mutagenesis studies, the reduced mobility of HIS516 and HIS559 observed in this work supports the notion that one of these histidine residues functions as the base catalyst in the overall catalytic cycle of GOx [27]. Collectively, these results indicate that the essential dynamic features of the GOx active site are preserved, maintaining a catalytically competent configuration that is consistent with previously reported native GOx systems.
To further characterize the structural stability and intermolecular interactions of the FAD@GOx systems, hydrogen bond analysis, radius of gyration (RoG), and solvent-accessible surface area (SASA) were evaluated over the 0-200 ns simulation period (Figure 4). The hydrogen bond analysis was performed to evaluate the stability of interactions between the ligand-receptor environment [59,60]. As shown in Figure 4A, the average number of hydrogen bonds involving the FAD cofactor remains relatively stable throughout the simulation period. The average number of hydrogen bonds involving the FAD cofactor in GOx was calculated over the 0-200 ns simulation period and summarized as follows: FAD@GOx (20.971 ± 0.026), Ce + FAD@GOx (23.429 ± 0.021), CS + FAD@GOx (27.155 ± 0.026), PVA + FAD@GOx (21.104 ± 0.023), and Ce + CS + PVA + FAD@GOx (18.059 ± 0.029). To further assess the persistence of these interactions, the hydrogen bond occupation percentages were calculated over the entire simulation timeframe. The results show high occupancy values for most systems, with FAD@GOx exhibiting 91.03% occupancy, followed by Ce + FAD@GOx (86.206%), CS + FAD@GOx (98%), PVA + FAD@GOx (38.583%), and Ce + CS + PVA + FAD@GOx (74.431%).
Various molecular dynamics-derived parameters of FAD@GOx systems: (A) Number of hydrogen bonds focusing on interactions between FAD and GOx, (B) RoG analyses for apo protein, and (C) SASA analysis active site for apo protein.
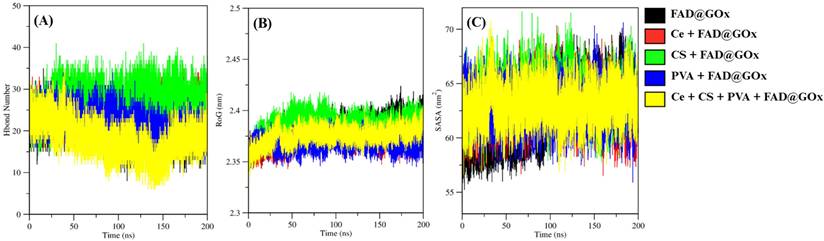
The radius of gyration (RoG) was used to evaluate the global compactness of the system during the simulations and surface accessible surface area (SASA) analysis reflect alterations in conformational stability and folding behavior throught the simulation time [61,62]. As shown in Figure 4B and Figure 4C, all systems exhibit stable RoG values and SASA throughout the 0-200 ns simulation period, indicating the absence of significant structural expansion or collapse. The average RoG values of the apoprotein in detailed: FAD@GOx (2.381 ± 0.000 nm), Ce + FAD@GOx (2.372 ± 0.000 nm), CS + FAD@GOx (2.388 ± 0.000 nm), PVA + FAD@GOx (2.367 ± 0.000 nm), Ce + CS + PVA + FAD@GOx (2.375 ± 0.000 nm), and the average SASA values of he apoprotein in detailed : FAD@GOx (60.230 ± 0.022 nm), Ce + FAD@GOx (62.994 ± 0.013 nm), CS + FAD@GOx (64.402 ± 0.013 nm), PVA + FAD@GOx (62.887 ± 0.012 nm), Ce + CS + PVA + FAD@GOx (63.203 ± 0.012 nm).
Per-residue energy decomposition was employed as a complementary analysis to support the interpretation of the overall binding free energy (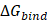 ), which serves as a key indicator of the interaction strength between FAD and GOx (Figure 5). In general,
), which serves as a key indicator of the interaction strength between FAD and GOx (Figure 5). In general, 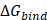 plays a crucial role in describing the thermodynamic favorability of ligand-receptor interactions, where more negative values correspond to spontaneous and more stonger binding process [63-65]. As shown in Table 2, the binding affinity follows the order FAD@GOx < PVA + FAD@GOx < Ce + CS + PVA + FAD@GOx < Ce + FAD@GOx < CS + FAD@GOx. The analysis further highlights specific amino acid residues that consistently contribute significantly to the binding stabilization, with individual energy contributions of ≤ -1.00 kcal·mol-1 [66], underscoring their dominant role in maintaining FAD anchoring within the GOx active site. Detailed contributions of the individual energy components are summarized in Table S3. It should be noted that, in the present molecular dynamics simulations, cerium was represented as a single Ce atom to simplify the computational model. While this approach provides preliminary insight into the interaction behavior, it may not fully capture the structural and electronic characteristics of CeO2 NPs. Therefore, the use of a more representative CeO2 NPs model could offer a more realistic description of atomic-level interactions within the system.
plays a crucial role in describing the thermodynamic favorability of ligand-receptor interactions, where more negative values correspond to spontaneous and more stonger binding process [63-65]. As shown in Table 2, the binding affinity follows the order FAD@GOx < PVA + FAD@GOx < Ce + CS + PVA + FAD@GOx < Ce + FAD@GOx < CS + FAD@GOx. The analysis further highlights specific amino acid residues that consistently contribute significantly to the binding stabilization, with individual energy contributions of ≤ -1.00 kcal·mol-1 [66], underscoring their dominant role in maintaining FAD anchoring within the GOx active site. Detailed contributions of the individual energy components are summarized in Table S3. It should be noted that, in the present molecular dynamics simulations, cerium was represented as a single Ce atom to simplify the computational model. While this approach provides preliminary insight into the interaction behavior, it may not fully capture the structural and electronic characteristics of CeO2 NPs. Therefore, the use of a more representative CeO2 NPs model could offer a more realistic description of atomic-level interactions within the system.
Peresidue energy decomposition profiles focus on interactions between FAD and GOx over the 125-150 ns; (A) FAD@GOx, (B) Ce + FAD@GOx, (C) CS + FAD@GOx, (D) PVA + FAD@GOx, and (E) Ce + CS + PVA + FAD@GOx.
Energy components (kcal mol-1) calculated using the MM-GBSA approach over the 125-150 ns. Values are reported as mean ± standard error of the mean (SEM), with a specific focus on interactions between FAD and GOx
| Component | ΔEvdw | ΔEele | ΔGgas | 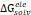 | 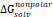 | 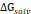 | 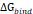 |
|---|---|---|---|---|---|---|---|
| FAD@GOx | -88.32 ± 0.13 | 78.61 ± 0.41 | -9.72 ± 0.47 | -67.32 ± 0.17 | -11.72 ± 0.00 | -79.04 ± 0.17 | -88.76 ± 0.50 |
| Ce + FAD@GOx | -87.31 ± 0.13 | 81.95 ± 0.55 | -5.36 ± 0.60 | -64.06 ± 0.02 | -11.37 ± 0.00 | -75.43 ± 0.02 | -80.79 ± 0.60 |
| CS + FAD@GOx | -82.14 ± 0.02 | 90.89 ± 0.27 | 8.75 ± 0.34 | -69.83 ± 0.46 | -10.95 ± 0.00 | -80.78 ± 0.46 | -72.03 ± 0.57 |
| PVA + FAD@GOx | -86.37 ± 0.14 | 83.19 ± 0.65 | -3.19 ± 0.70 | -70.37 ± 0.30 | -11.53 ± 0.00 | -81.90 ± 0.30 | -85.08 ± 0.76 |
| Ce + CS + PVA + FAD@GOx | -83.88 ± 0.15 | 73.01 ± 0.51 | -10.87 ± 0.55 | -61.09 ± 0.20 | -11.16 ± 0.00 | -72.24 ± 0.20 | -83.11 ± 0.59 |
Ultraviolet visible spectrophotometry (UV-Vis) study
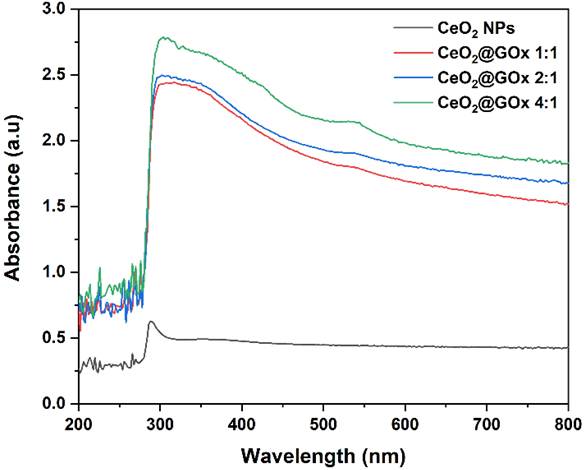
The denaturation of GOx upon adsorption onto nanostructured surfaces is generally associated with alterations in its molecular conformation, which may lead to partial or complete loss of enzymatic activity [67]. Thus, UV-Vis spectroscopy was utilized to monitor potential alterations in the electronic and conformational structure of GOx during the modification process. UV-Vis was used to examine the possible conformation changes of GOx following the incorporation of CeO2 NPs. As shown in Figure 6, UV-Vis analysis shows a peak at a wavelength of 296 nm which indicates the presence of absorbance in CeO2 NPs [68,69]. Furthermore, the presence of native GOx is indicated by an absorption band at ~380 nm, which is attributed to the polypeptide chains, while a slight red-shift toward ~530 nm can be associated with the flavin prosthetic group within the protein structure [70,67]. Increasing the CeO2@GOx ratio from 1:1, 2:1, to 4:1 causes an increase in absorbance which indicates the concentration of CeO2 NPs [37,71].
UV-Vis characterization spectrum of CeO2@GOx.
Fourier transform infrared spectroscopy (FT-IR) study
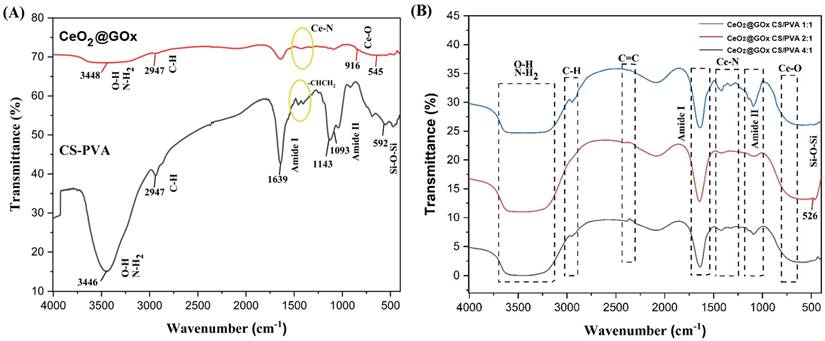
Fourier transform infrared (FT-IR) characterization was performed on CeO2@GOx CS/PVA hydrogels with mass ratios of 1:1, 2:1, and 4:1 in the range of 4000-500 cm-1. The FT-IR spectrum of CS/PVA exhibits broad absorption peaks at 3000-3700 cm-1, attributed to O-H and N-H2 stretching vibrations, along with C-H stretching vibrations at 2947 cm-1. The characteristic sharp peaks belonging to amide I and amide II bands, as well as bending vibrations corresponding to -CHCH2 (Figure 7A). The spectrum of CeO2@GOx shows a characteristic metal-oxygen stretching vibrations (Ce-O) at ~535 cm-1, while the band at 1035 cm-1 represented as O-Ce-O bond [72,73]. As shown in Figure 7B, a weak band observed at 1000-1500 cm-1 occurs due to the complexation between Ce and amine groups in that area [74].
FT-IR characterization results; (A) Ce@GOx and CS/PVA; (B) Hydrogel variations of Ce@GOx CS/PVA 1:1, Ce@GOx CS/PVA 2:1, and Ce@GOx CS/PVA 4:1.
Particle size analyzer (PSA) study
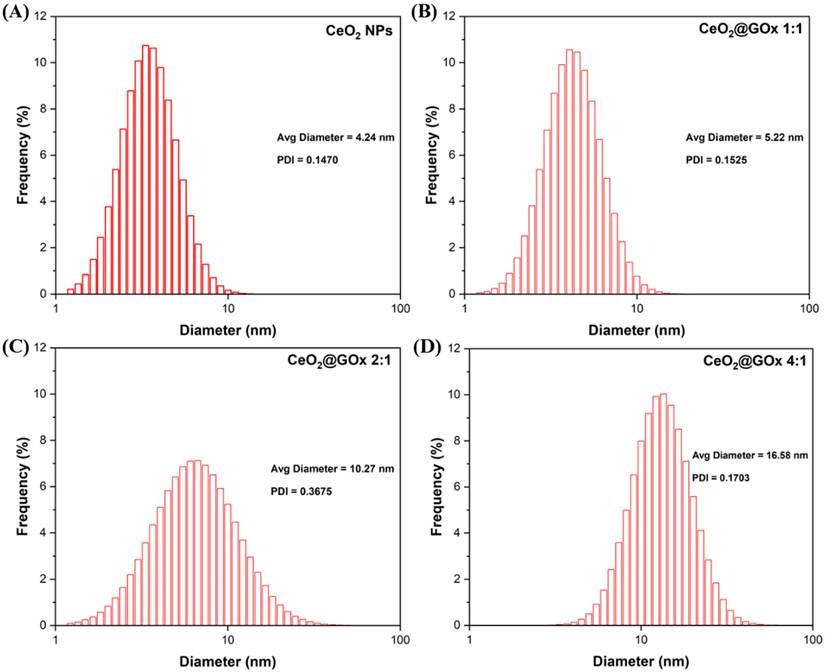
Characterization using Particle Size Analyzer (PSA) was performed to determine the range and distribution of particle size especially after the synthesis process of CeO2 NPs and functionalization of CeO2 NPs with GOx based on the principle of Brownian motion. According to the characterization with a measurement range of 1nm-1000nm, the average diameter of the CeO2 NPs sample (Figure 8A) was 7.76 nm with a Polydispersity Index (PDI) of 0.3668 indicating the successful formation of CeO2 NPs. The Ce@GOx 1:1 sample (Figure 8B) has an average diameter of 5.22 nm with a PDI of 0.1525. The CeO2@GOx 2:1 sample (Figure 8C) has an average diameter of 10.27nm with a PDI of 0.3675. The CeO2@GOx 4:1 sample (Figure 8D) has an average diameter of 16.58nm with a PDI of 0.1703. The CeO2@GOx 2:1 and CeO2@GOx 4:1 samples showed an average particle size that was in accordance with the recomended particle size range for transdermal route drug administration of 10nm-600nm [75]. In addition, all four samples have a PDI value ≤ 0.3 which indicates a homogeneous particle size distribution and is acceptable for industrial drug applications [76].
PSA characterization results; (A) CeO2 NPs, (B) CeO2@GOx 1:1, (C) CeO2@GOx 2:1, and (D) CeO2@GOx 4:1.
Degradation test
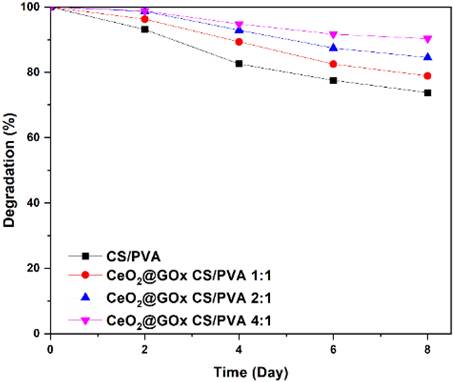
The weight loss of the hydrogels was monitored in phosphate buffer saline (PBS) at room temperature over time. The hydrogel weight decreased significantly up to day 8, as shown in (Figure 9). CeO2@GOx CS/PVA 4:1 hydrogel resulted in degradation with the highest mass reduction of 90.3%. The degradation rate of CeO2@GOx CS/PVA 2:1 hydrogel was slightly lower, with approximately 84.575% of the initial weight remaining at day 8. The degradation rate of CeO2@GOx CS/PVA 1:1 hydrogel remained at 78.9%. Meanwhile, the CS/PVA hydrogel showed the lowest mass reduction, which was 73.7%. This mass reduction is due to the breakdown of the crosslinking bonds that hold the hydrogel structure [13]. Degradation performance is very important in applying materials as wound healing in wounds, especially in diabetic wounds [77]. The application of hydrogels in wound healing must have high physical resistance with low degradation rates to maintain their structure and function in cell and tissue regeneration for a certain period of time.
Degradation test of the CeO2@GOx hydrogel sample in CS/PVA for every variation.
Enzymatic activity (peroxidase and catalase)
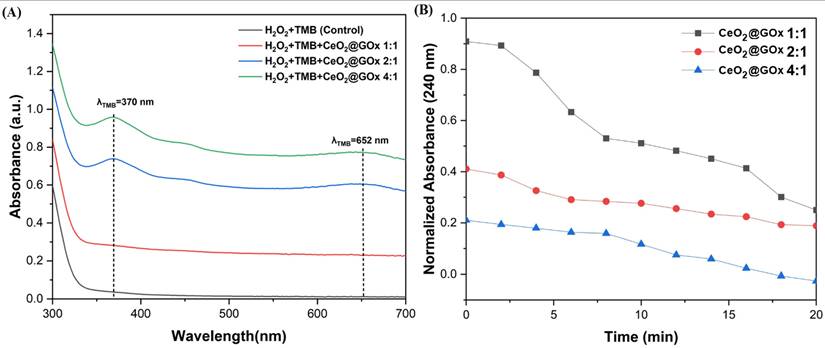
The enzymatic peroxidase activity of CeO2@GOx sample was tested through the oxidation rate of 3,3′,5,5′-tetramethylbenzidine (TMB). As shown in Figure 10A, the sample exhibited good affinity towards TMB in the presence of H2O2, as evidenced by the appearance of dual characteristic peaks at 370 nm and 652 nm, which are indicative of typical nanozyme behavior [78,79]. Furthermore, the catalase-like activity of CeO2@GOx was demonstrated by the concentration-dependent degradation of H2O2, monitored via the decrease in absorbance at 240 nm (Figure 10B) [80]. According to previous studies, CeO2 is capable of catalytically decomposing H2O2 to regenerate oxygen, thereby protecting GOx from oxidative denaturation [81]. In the CeO₂@GOx system, glucose oxidation proceeds through the canonical catalytic cycle of glucose oxidase (GOx). According to Ernst et al. [82], glucose molecules are oxidized by GOx in its flavin adenine dinucleotide (FAD) form, producing gluconolactone and reducing GOx(FAD) to GOx(FADH₂). The gluconolactone intermediate subsequently hydrolyzes to form gluconic acid. The reduced enzyme is then re-oxidized by molecular oxygen, leading to the generation of hydrogen peroxide (H₂O₂) as a key reaction intermediate. In the presence of CeO₂, the locally generated H₂O₂ can be dynamically regulated through its peroxidase- and catalase-like activities, thereby mitigating oxidative stress and preserving the structural integrity of GOx. Here, the catalytic cycle of GOx is based on previous research [82,83]:
Enzymatic activity analysis; (A) Peroxidase, and (B) Catalase.
GOx(FAD) + β-D-glucose → GOx(FADH₂) + gluconolactone (11)
GOx(FADH₂) + O2 → GOx(FAD) + H2O2 (12)
H2O2 → O2 + 2e- + 2H+ (13)
Antimicrobial assay
Disc diffusion assay (DDA) results for CeO2@GOx sample variations and an antimicrobial agent named gentamicin against the growth of Gram-negative bacteria Escherichia coli ATCC 25922 and Gram-negative bacteria Staphylococcus aureus ATCC 25923 are presented in Table 3 and Table 4. The results indicate moderate antimicrobial activity from CeO2@GOx sample variations with all average inhibition zones not lower than 6 mm [84,85]. Yet the results are still far from a commercial antibacterial agent such as gentamicin, which shows a strong antimicrobial activity. In addition, CeO2 NPs can bind to microbial proteins and disrupt essential functions. The released cerium ions interfere with respiration and electron flow in microbial cells. CeO2 NPs are also known to interact with thiol groups (-SH) and membrane transporters or porins, hindering the transport of nutrients into the cell. Another critical antimicrobial mechanism is oxidative stress [86,87]. The reversible cycling between Ce(III) and Ce(IV) states on the surface of microbial membranes leads to the generation of reactive oxygen species (ROS). These ROS attack vital cellular components such as lipids, proteins, polysaccharides, and nucleic acids, resulting in functional damage and eventual microbial cell death. Although the CeO2@GOx CS/PVA nanocomposite demonstrated antimicrobial activity, the observed effects were categorized as moderate. This outcome suggests several possible limitations that warrant further evaluation. First, the enzymatic activity of glucose oxidase (GOx) might have been partially compromised during nanoparticle functionalization or matrix incorporation, potentially reducing hydrogen peroxide (H₂O₂) generation. The limited production of H₂O₂ would weaken the oxidative stress required for effective microbial inhibition. Additionally, the redox capacity of cerium oxide nanoparticles depends heavily on the Ce(III)/Ce(IV) ratio, which may not have been optimized, thus diminishing reactive oxygen species (ROS) generation [88,89].
Inhibition zone diameter against the growth of Escherichia coli ATCC 25922
| No. | Sample Code | Inhibition Zone Diameter (mm) | Average Inhibition Zone (mm) | ||
|---|---|---|---|---|---|
| D1 | D2 | D3 | |||
| 1 | K(+)* | 20.35 | 19.86 | 19.90 | 20.10 |
| 2 | K(-)* | 0 | 0 | 0 | 0 |
| 3 | CeO2@GOx 1:1 | 7.36 | 7.07 | 7.63 | 7.35 |
| 4 | CeO2@GOx 2:1 | 6.90 | 6.81 | 6.71 | 6.81 |
| 5 | CeO2@GOx 4:1 | 6.45 | 6.44 | 6.71 | 6.54 |
*K(+) : Gentamicin K(-) : Aquades
Inhibition zone diameter against the growth of Staphylococcus aureus ATCC 25923
| No. | Sample Code | Inhibition Zone Diameter (mm) | Average Inhibition Zone (mm) | ||
|---|---|---|---|---|---|
| D1 | D2 | D3 | |||
| 1 | K(+)* | 30.65 | 29.73 | 24.55 | 30.19 |
| 2 | K(-)* | 0 | 0 | 0 | 0 |
| 3 | CeO2@GOx 1:1 | 7.97 | 8.63 | 8.38 | 8.33 |
| 4 | CeO2@GOx 2:1 | 7.59 | 7.88 | 7.72 | 7.73 |
| 5 | CeO2@GOx 4:1 | 7.85 | 8.02 | 8.97 | 8.28 |
*K(+) : Gentamicin K(-) : Aquades
Another contributing factor could be the matrix composition. While chitosan and PVA provide structural and biocompatible support, their interaction with CeO2 NPs and GOx may restrict the diffusion or release of active agents. A high ratio of PVA, for instance, could create a dense polymer network that hinders effective contact with microbial cells. Furthermore, the release profile of H₂O₂ or Ce ions may be too slow to induce significant antimicrobial effects within the test duration. Environmental factors during testing, such as the pH or availability of glucose, may also have played a role. GOx activity is substrate-dependent; thus, insufficient glucose concentration in the medium would limit the enzymatic reaction and subsequent H₂O₂ production. The type and resistance level of the tested microbial strains could further influence the outcome, especially if the bacteria possess tolerance mechanisms against oxidative stress. To improve antimicrobial performance, future work should consider optimizing the GOx and CeO2 NPs loading ratios, adjusting the CS/PVA composition, and confirming enzymatic and redox activities through H₂O₂ and ROS quantification assays. Additionally, pre-incubation strategies or combining this system with other antimicrobial agents may offer synergistic effects to enhance overall efficacy.
In vivo test results
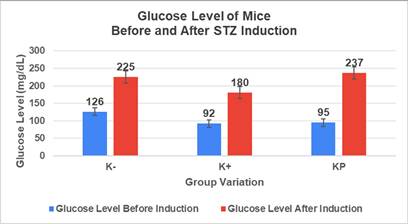
The results showed a significant increase in blood glucose levels in various treatment groups after STZ induction. Mice are declared diabetic if blood glucose levels exceed 175 mg/dL [90,91]. Based on these criteria and Figure 13, it can be concluded that the administration of STZ is effective in inducing diabetic conditions in mice, with blood glucose levels in all groups showing hyperglycemia for 5 days. Visual observation results in (Table 5) show KP as the group with wound healing rate close to KN, indicating the use of CeO2@GOx CS/PVA 4:1 hydrogel as a sample with superior healing effect compared to the use of conventional drugs. Nevertheless, comprehensive histopathological analysis is required to further validate the quality of tissue regeneration, modulation of the inflammatory response, and collagen remodeling at the microscopic level. Such investigations will provide deeper insight into the therapeutic mechanism and further strengthen the clinical potential of the developed hydrogel system.
Zone of inhibition test results against the growth of Escherichia coli ATCC 25922; (A) First-replication, (B) Second-replication, and (C) Third-replication.
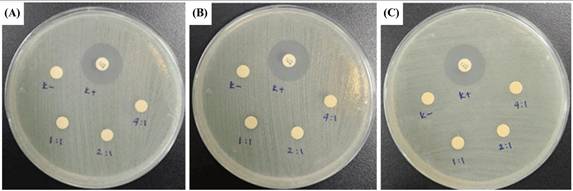
Zone of inhibition test results against the growth of Staphylococcus aureus ATCC 25923; (A) First-replication, (B) Second-replication, and (C) Third-replication.
Diabetes induction in mice.
Results of visual observation of diabetic wound healing in mice
| Treatment group | Day 1 | Day 2 | Day 3 | Description |
|---|---|---|---|---|
| KN |  | 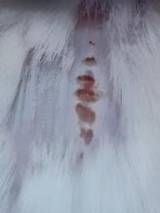 |  | Non-diabetic, wounded, and untreated. Mice showed no inflammation and proliferation was rapid until the wound was completely closed by day 7. |
| K- | 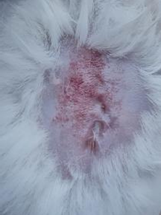 | 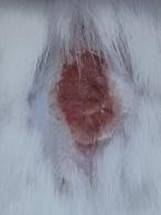 | 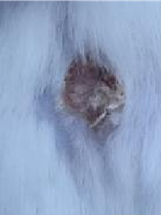 | Diabetic, wounded, and untreated. The mice showed prolonged inflammation and proliferation until the wound had not entered the maturation stage by day 7. |
| K+ |  |  | 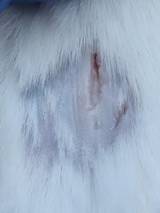 | Diabetics, wounded, and administered 10% betadine ointment (Transdermal). The mice showed no inflammation and moderate proliferation until the 7th day when the wound entered the maturation stage. |
| KP | 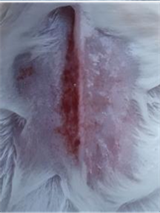 |  |  | Diabetics, wounded, and administered CeO2@GOx CS/PVA composite hydrogel variation 4:1. The mice showed no inflammation and proliferation was rapid until the wound was completely closed on the 7th day. |
Conclusion
CeO2@GOx hydrogels in CS/PVA solution were successfully synthesized through four main stages, namely CeO2 NPs synthesis, CS/PVA hydrogel formation, CeO2@GOx incorporation, and composite hydrogel formation by freeze-thawing method. Characterization analysis showed that increasing the CeO2@GOx ratio from 1:1, 2:1, to 4:1 led to an increase in absorbance, indicating an increasing Ce concentration. Degradation and enzymatic activity tests showed that CeO2@GOx CS/PVA hydrogels have good potential as wound healing materials. In vivo tests on mice with diabetic wound models showed that the use of CeO2@GOx 4:1 CS/PVA hydrogel provided a better healing effect compared to conventional drugs, with wounds completely closed on day 7. The results of this study indicate that CeO2@GOx CS/PVA hydrogel has potential as an innovation in diabetic wound management.
Supplementary Material
Supplementary figures and tables.
Acknowledgements
The authors would like to thank the National Research and Innovation Agency (BRIN) for providing access to computational facilities.
Funding
This research is funded by the Indonesian Endowment Fund for Education (LPDP) on behalf of the Indonesian Ministry of Higher Education, Science, and Technology and managed under the EQUITY Program (Contract No. 4300/B3/DT.03.08/2025; No. 297/UN3/HK.07.00/2025) and Universitas Airlangga following contract (5554/B/UN3.LPPM/PT.01.03/2025).
Author contributions
S.H. led the project, designed the experiments, analysed the particle characterization experiments, wrote the manuscript; L.G.N synthesis the material, performed characterization, in vivo test, antimicrobial assay, and wrote the manuscript; D.P.P and R.A.P synthesis the material and the hydrogel, performed characterization, and performed the in silico analysis; A.M.K, D.S.T.S performed the enzymatic test, antibacterial analysis, and in vivo analysis; M.I.B and W.H provided support and guidance on this project, performed data analysis on antibacterial test; N.A.N.N.M provided guidance and expertise on material characterization and revised the manuscript; I.S performed in silico analysis and revised the manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Winarni D, Husna FN, Syadzha MF, Susilo RJK, Hayaza S, Ansori ANM, Alamsjah MA, Amin MNG, Wulandari PAC, Pudjiastuti P, Awang K. Topical Administration Effect of Sargassum duplicatum and Garcinia mangostana Extracts Combination on Open Wound Healing Process in Diabetic Mice. Scientifica. 2022 1, p. 9700794
2. Kharroubi AT, Darwish HM. Diabetes mellitus: The epidemic of the century. World Journal of Diabetes. 2015;6:6 pp. 850-867
3. Muchtaromah B, Firdaus AMK, Ansori ANM, Duhita MR, Minarno EB, Hayati A, Ahmad M, Analisa I. Effect of pegagan (Centella asiatica) nanoparticle coated with chitosan on the cytokine profile of chronic diabetic mice. Narra J. 2024: 4, 1, pp. e697-e697.
4. Marbun PIG, Mardiani TH. Correlation Between Blood Glucose Level And Thinking Concentration. FMI. 2016;52:3
5. Lede MJ, Hariyanto T, Ardiyani VM. Pengaruh Kadar Gula Darah Terhadap Penyembuhan Luka Diabetes Mellitus Di Puskesmas Dinoyo Malang. Nursing News : Jurnal Ilmiah Keperawatan. 2018;3:1
6. Sathish GPBS, Joseph LMIL, Subramanian KM, Kalist V, Vasanth K. Retinal Blood Vessels and Optical Disc Segmentation in Branch Retinal Vein Occluded Fundus Images Using Digital Image Processing Techniques. Research Journal of Pharmacy and Technology. 2019 12:, 4, pp. 1901-1906
7. Odularu AT, Ajibade PA. Challenge of diabetes mellitus and researchers' contributions to its control. Open Chemistry. 2021;19:1 pp. 614-634
8. Aynalem SB, Zeleke AJ. Prevalence of Diabetes Mellitus and Its Risk Factors among Individuals Aged 15 Years and Above in Mizan-Aman Town, Southwest Ethiopia, 2016: A Cross Sectional Study. International Journal of Endocrinology. 2018: 1, p. 9317987.
9. Hidayatillah S, Heri N, Sakundarno M. Hubungan Status Merokok dengan Kejadian Ulkus Diabetikum pada Laki-Laki Penderita Diabetes Melitus. Jurnal Epidemiologi Kesehatan Komunitas. 2020;5:pp 32-37
10. Zainuddin SZ, Ramond NJMR, Anuar NK. Polysaccharide-Based Formulations for the Treatment of Diabetic Wounds: A Review. Research Journal of Pharmacy and Technology. 2023;16:6 pp. 2835-2842
11. Xu C, Qu X. Cerium oxide nanoparticle: a remarkably versatile rare earth nanomaterial for biological applications. NPG Asia Materials. 2014;6:3 pp. e90-e90
12. Zhu Y, Zhang J, Song J, Yang J, Dua Z, Zhao W, Guo H, Wen C, Li Q, Sui X, Zhang L. A Multifunctional Pro-Healing Zwitterionic Hydrogel for Simultaneous Optical Monitoring of pH and Glucose in Diabetic Wound Treatment. Advanced Functional Materials. 2020;30:6 p. 1905493
13. Mohammed ZMA, Baydaa J, Hassan A. The effect of Glucoamylase and Glucose oxidase from the Aspergillus niger and Penicillium notatum respectively on inhibition of Streptococcus mutans causes the Dental caries. Research Journal of Pharmacy and Technology. 2018;11:8 pp. 3419-3422
14. Tsuruoka N, Soto SS, Tahar AB, Zebda A, Tsujimura S. Mediated electrochemical oxidation of glucose via poly(methylene green) grafted on the carbon surface catalyzed by flavin adenine dinucleotide-dependent glucose dehydrogenase. Colloids and Surfaces B: Biointerfaces. 2020;192:p 111065
15. Waly Y, Hussain A, Al-Majmuei A, Alatoom M, Alaraibi AJ, Alaysereen A, Deen GR. Evolution of Biosensors and Current State-of-the-Art Applications in Diabetes Control. Biosensors. 2026;16:1 p. 39
16. Lee SH, Choi DS, Kuk SK, Park CB. Photobiocatalysis: Activating Redox Enzymes by Direct or Indirect Transfer of Photoinduced Electrons. Angewandte Chemie International Edition. 2018;57:27 pp. 7958-7985
17. He L, Lin Z, Zou H, Liu Q, Liu Y, Liu H, Wu X, Pan X. Glucose oxidase-triggered cascade reaction in nanocomposite hydrogel for on-demand nitric oxide delivery and diabetic wound regeneration. International Journal of Biological Macromolecules. 2026;341:p 150127
18. Neufeld L, Bianco-Peled H. Pectin-chitosan physical hydrogels as potential drug delivery vehicles. International Journal of Biological Macromolecules. 2017;101:pp 852-861
19. D S, R S, R L. et al. Synthesis and Characterization of MgO nanoparticles Coated by polyvinyl alcohol for Biomedical applications. Research Journal of Pharmacy and Technology. 2018;11:pp 4997-5000
20. Latha AM, Ravikiran TN, Kumar JNS. Formulation, Optimization and Evaluation of Glibenclamide Transdermal Patches by using chitosan Polymer. Asian Journal of Pharmacy and Technology. 2019;9:1 pp. 01-07
21. Bertoli AC, Novaes PAA, Costa LT, Almeida WBD, Duarte HA. Insights into rare earth element speciation: unraveling sulfate and hydrolysis complexes through DFT calculations. J Mol Model. 2024;30:12 p. 412
22. Pan X, Wu C, Fang H, Yan C. Structural, Spectroscopic, and Bonding Analyses of La(III)/Ce(III)-Tetrel Ate-Complexes. Inorganic Chemistry. 2023;62:14 pp. 5660-5668
23. Guo R, Wang L, Cai Z, Zhao Z, Bian Z, Liu Z. Complexes of Ce(III) and Bis(pyrazolyl)borate Ligands: Synthesis, Structures, and Luminescence Properties. Inorganic Chemistry. 2022;61:35 pp. 14164-14172
24. Sachin AR, Gopakumar G, Rao CVSB. Understanding the Complexation Behavior of Carbamoylphosphine Oxide Ligands with Representative f-Block Elements. The Journal of Physical Chemistry A. 2024;128:6 pp. 1085-1097
25. Tomeček J, Liddle ST, Kaltsoyannis N. Computational Investigations of Metal-Metal Bonding in Molecular Thorium Compounds and Ce and Group IV Analogues. Inorganic Chemistry. 2025;64:28 pp. 14498-14512
26. Shi J, Ren Q, Gao Y. Accelerating Global Optimization of Cerium Oxide Nanocluster Structures with High-Dimensional Neural Network Potential. The Journal of Physical Chemistry A. 2025;129:9 pp. 2190-2199
27. Puspita PJ, Ambarsari L, Adiva A, Sumaryada TI. In silico Analysis of Glucose Oxidase H516r and H516d Mutations for an Enzymatic Fuel Cell. Jurnal Kimia Valensi. 2021: pp. 83-93.
28. Sari WON, Hardjo M, Syahrijuita, Azis I, Yustisia I. In silico Evaluation of 3,6-Anhydro-D-Galactose from Eucheuma denticulatum Targeting Breast Cancer-Associated Molecular Pathways. Tropical Journal of Natural Product Research. 2025;9:11 p. 5497
29. Gowsalya R, P R, M S. Eco-friendly cerium vanadate nanoparticles for UTI control: integrating molecular docking and antibacterial evaluation. Physica Scripta. 2025;100:8 p. 085915
30. Janoš P, Henych J, Pfeifer J, Zemanová N, Pilařová V, Milde D, Opletal T, Tolasz J, Malýd M, Štengl V. Nanocrystalline cerium oxide prepared from a carbonate precursor and its ability to breakdown biologically relevant organophosphates. Environmental Science: Nano. 2017;4:6 pp. 1283-1293
31. Tu T, Zhang Y, Yan Y, Li L, Liu X, Hakulinen N, Zhang W, Mu Y, Luo H, Yao B, Li W, Huang H. Revealing the intricate mechanism governing the pH-dependent activity of a quintessential representative of flavoproteins, glucose oxidase. Fundamental Research. 2024
32. Verma G, Hostert J, Summerville A, Robang AS, Carcamo RG, Paravastu AK, Getman RB, Duval CE, Renner J. Investigation of Rare Earth Element Binding to a Surface-Bound Affinity Peptide Derived from EF-Hand Loop I of Lanmodulin. ACS Applied Materials & Interfaces. 2024;16:13 pp. 16912-16926
33. Verma AK, Sharma S, Jayaraj A, Deep S. In silico study of interaction of (ZnO)12 nanocluster to glucose oxidase-FAD in absence and presence of glucose. Journal of Biomolecular Structure and Dynamics. 2023;41:no 24, pp. 15234-15242
34. Ekasurya W, Sebastian J, Puspitasari D, Asri PPP, Asri LATW. Synthesis and Degradation Properties of Sericin/PVA Hydrogels. Gels. 2023;9:2 Art. no. 2
35. Singh S. Cerium oxide based nanozymes: Redox phenomenon at biointerfaces. Biointerphases. 2016;11:no 4, p. 04B202
36. George G, Wilson R, Joy J. Chapter 3 - Ultraviolet Spectroscopy: A Facile Approach for the Characterization of Nanomaterials. In Spectroscopic Methods for Nanomaterials Characterization. 2017: pp. 55-72.
37. Yu X, Fu X, Yang J. Glucose/ROS cascade-responsive ceria nanozymes for diabetic wound healing. Materials Today Bio. 2022;15:p 100308
38. Liu Z, Wan P, Yang M, Han F, Wang T, Wang Y, Li Y. Cell membrane camouflaged cerium oxide nanocubes for targeting enhanced tumor-selective therapy. Journal of Materials Chemistry B. 2021;9:46 pp. 9524-9532
39. Nene R, Pandey S, Gidwani B, Pandey RK, Shukla SS. Fourier Transform Infra-Red Spectroscopy: Recent advances and prospective in Analytical Method Development and Validation. Research Journal of Pharmacy and Technology. 2024;17:6 pp. 2955-2958
40. Dhanaraj S, Bharathiraja P, Dhandapani R, Subbaiya R, Kahiresha AK. Biosynthesis and Characterization of Silver Nanoparticles from Aspergillus niger and its Antibacterial Activity. Research Journal of Pharmacy and Technology. 2018;11:12 pp. 5282-5286
41. Babick F. Chapter 3.2.1 - Dynamic light scattering (DLS) in Characterization of Nanoparticles. Micro and Nano Technologies. 2020: pp. 137-172
42. Hanif W, Hardiansyah A, Randy A, Asri LATW. Physically crosslinked PVA/graphene-based materials/aloe vera hydrogel with antibacterial activity. RSC Advances. 2021;11:46 pp. 29029-29041
43. Bhagat S, Vallabani NVS, Shutthanandan V, Bowden M, Karakoti AS, Singh S. Gold core/ceria shell-based redox active nanozyme mimicking the biological multienzyme complex phenomenon. Journal of Colloid and Interface Science. 2018;513:pp 831-842
44. Mohite SA, Shah RR, Patel NR. Antimicrobial Activity of Leaves extracts of Jatropha curcas. Asian Journal of Pharmaceutical Research. 2018;8:1 pp. 17-20
45. Purohit MC, Kandwal A, Purohit R, Semwal AR, Parveen S, Khajuria AK. Antimicrobial Activity of Synthesized Zinc Oxide Nanoparticles using Ajuga bracteosa Leaf Extract. Asian Journal of Pharmaceutical Analysis. 2021;11:4 pp. 275-280
46. Miar M, Shiroudi A, Pourshamsian K, Oliaey AR, Hatamjafari F. Theoretical investigations on the HOMO-LUMO gap and global reactivity descriptor studies, natural bond orbital, and nucleus-independent chemical shifts analyses of 3-phenylbenzo[d]thiazole-2(3H)-imine and its para-substituted derivatives: Solvent and substituent effects. Journal of Chemical Research. 2021;45:1-2 pp. 147-158
47. Hussein HH, Fadhil GF. Investigation of structural and conformational stability, electronic transition, NLO, FMO, and DSSC parameters of trans-dichloro-nitro chalcone isomers: a DFT insight. Journal of Computational Electronics. 2025;24:5 p. 140
48. Krishnasamy K, Subramani T. Unveiling the molecular mechanism of acyclovir interaction with carbon dots: a DFT approach. Journal of Molecular Modeling. 2025;32:1 p. 6
49. Mondal A, Srivastava A, Acharya K, Boruah PJ, Chamlagai D, Kamra A, Bhattacharya A, Rana S, Roy AS. Dual-functional aggregation-induced emission active chitosan-based hydrogel for the detection and killing of pathogenic bacteria. Biomaterials Science. 2025
50. Ixtiyorovich MF. Study Of The Electronic Structure Of The Vanadyl (Ii) Monochloroacetate Monomer Molecule. Nvpubhouse Library for American Journal of Applied Science and Technology. 2025;5:09 pp. 63-67
51. Lü R, Lin J, Zhao X, Zhao X. Theoretical Study on Interactions Between Ionic Liquid and Chitin/Chitosan/Cellulose. Journal of the Chilean Chemical Society. 2017;62:4 pp. 3668-3676
52. Purvis GD, Culberson C. On the graphical display of molecular electrostatic force-fields and gradients of the electron density. Journal of Molecular Graphics. 1986;4:2 pp. 88-92
53. Morales-Pumarino D, Barquera-Lozada JE. Electron density and its reduced density gradient in the study of π-π interactions,” International Journal of Quantum Chemistry. 2023; 123: 18, p. e27051.
54. Manu P, Abakah A, Anfu PK, Osei-Poku P, Kwarteng A. Disrupting quorum sensing by exploring conformational dynamics and active site flexibility of LasR protein in Pseudomonas aeruginosa. Discovery Chemistry. 2025;2:1 p. 39
55. Adeniran AA, Adenaya A, Ogunlakin AD, Prabhu D, Sonibare MA. Quercetin-3-O-galactoside from Garcinia Kola Heckel stem bark as a potential bioactive inhibitor against histone deacetylase 8 (HDAC8) using in silico study. Discovery Chemistry. 2025;2:1 p. 304
56. Wang F, Chen X, Wang Y, Li X, Wan M, Zhang G, Leng F, Zhang H. Insights into the Structures, Inhibitors, and Improvement Strategies of Glucose Oxidase. International Journal of Molecular Sciences. 2022;23:17 p. 9841
57. Kornecki JF, Carballares D, Tardioli P, Rodrigues RC, Berenguer-Murcia Ángel Alcántarae AR, Fernandez-Lafuente R. Enzyme production of D-gluconic acid and glucose oxidase: successful tales of cascade reactions. Catalysis Science & Technology. 2020;10:17 pp. 5740-5771
58. Witt S, Wohlfahrt G, Schomburg D, Hecht HJ, Kalisz HM. Conserved arginine-516 of Penicillium amagasakiense glucose oxidase is essential for the efficient binding of β-d-glucose. Biochemical Journal. 2000;347:2 pp. 553-559
59. Huang YM, Kang M, Chang C. A. Switches of hydrogen bonds during ligand-protein association processes determine binding kinetics. Journal of Molecular Recognition. 2014;27:9 pp. 537-548
60. Blazhynska M, Ansari N, Lagardère L, Piquemal J-P. From Water Networks to Binding Affinities: Resolving Solvation Dynamics for Accurate Protein-Ligand Predictions. bioRxiv. 2025
61. Shankar U, Jain N, Majee P, Kodgire P, Sharma TK, Kumar A. Exploring Computational and Biophysical Tools to Study the Presence of G-Quadruplex Structures: A Promising Therapeutic Solution for Drug-Resistant Vibrio cholerae. Frontiers in Genetics. 2020: 11, Sep. 2020
62. Sharma K, Fontes CMGA, Najmudin S, Goyal A. Small angle X-ray scattering based structure, modeling and molecular dynamics analyses of family 43 glycoside hydrolase α-L-arabinofuranosidase from Clostridium thermocellum. Journal of Biomolecular Structure and Dynamics. 2021;39:1 pp. 209-218
63. Thakur A, Patwa J, Pant S, Sharma A, Flora SJS. Interaction study of monoisoamyl dimercaptosuccinic acid with bovine serum albumin using biophysical and molecular docking approaches. Scientific Reports. 2021: 11: 1, p. 4068.
64. Choudhary M. I. Shaikh M, -tul-Wahab A, -ur-Rahman A. In silico identification of potential inhibitors of key SARS-CoV-2 3CL hydrolase (Mpro) via molecular docking, MMGBSA predictive binding energy calculations, and molecular dynamics simulation. PLOS ONE. 2020;15:7 p. e0235030
65. Jusoh N, Zainal H, Hamid AAA, Bunnori NM, Halim KBA, Hamid SA. In silico study of carvone derivatives as potential neuraminidase inhibitors. Journal of Molecular Modeling. 2018;24:4 p. 93
66. Kristanti AN, Aminaha NS, Siswanto I, Wardana AP, Abdjan MI, Khoirunisak AR, Noviana E. Synthesis, Pharmacokinetic, Molecular Docking, and Molecular Dynamics Simulation of 2-Styrylchromone Derivatives as Potential Inhibitor of Human Kinesin Eg5. Engineered Science. 2024;30:1168
67. Zhang X, Liu D, Li L, You T. Direct Electrochemistry of Glucose Oxidase on Novel Free-Standing Nitrogen-Doped Carbon Nanospheres@Carbon Nanofibers Composite Film. Scientific Reports. 2015;5:1 p. 9885
68. Madkour M, Allam OG, Nazeer AA, Amin MO, Al-Hetlani E. CeO2-based nanoheterostructures with p-n and n-n heterojunction arrangements for enhancing the solar-driven photodegradation of rhodamine 6G dye. Journal of Materials Science: Materials in Electronics. 2019;30:11 pp. 10857-10866
69. Sozarukova MM, Koziova TO, Beshkareva TS, Popov AL, Kolmanovich DD, Vinnik DA, Ivanova OS, Lukashin AV, Baranchikov AE, Ivanov VK. Gadolinium Doping Modulates the Enzyme-like Activity and Radical-Scavenging Properties of CeO2 Nanoparticles. Nanomaterials. 2024;14:9 p. 769
70. Xu X, Wu P, Xu W, Shao Q, An L, Zhang H, Cai C, Zhao B. Effects of guanidinium ions on the conformational structure of glucose oxidase studied by electrochemistry, spectroscopy, and theoretical calculations: towards developing a chemical-induced protein conformation assay. Physical Chemistry Chemical Physics. 2012;14:16 pp. 5824-5836
71. Singh R, Singh S. Redox-dependent catalase mimetic cerium oxide-based nanozyme protect human hepatic cells from 3-AT induced acatalasemia. Colloids and Surfaces B: Biointerfaces. 2019;175:pp 625-635
72. Sana SS, Mishra V, Vadde R, Silanpaa Mika, Alfarraj S, Hung PV, Kim S-C. Curcuma longa debranched starch assisted synthesis of cerium oxide nanoparticles and its antioxidant, anticancer, antimicrobial, and anti-biofilm activities. Scientific Reports. 2026
73. Hkiri K, Mohamed HEA, Ghotekar S, Maaza M. Green synthesis of cerium oxide nanoparticles using Portulaca oleracea Extract: Photocatalytic activities. Inorganic Chemistry Communications. 2024;162:p 112243
74. Kalantari K, Mostafavi E, Saleh B, Soltantabar P, Webster TJ. Chitosan/PVA hydrogels incorporated with green synthesized cerium oxide nanoparticles for wound healing applications. European Polymer Journal. 2020;134:p 109853
75. Danaei M, Dehghankhold M, Ataei S, Davarani FH, Javanmard R, Dokhani A, Khorasani S, Mozafari MR. Impact of Particle Size and Polydispersity Index on the Clinical Applications of Lipidic Nanocarrier Systems. Pharmaceutics. 2018;10:2 p. 57
76. Chen M, Liu X, Fahr A. Skin penetration and deposition of carboxyfluorescein and temoporfin from different lipid vesicular systems: In vitro study with finite and infinite dosage application. International Journal of Pharmaceutics. 2011;408:1 pp. 223-234
77. Kuppusamy KCG, Saikamal, Karri VVSR. Simvastatin Loaded Polycaprolactone-collagen Scaffolds for the treatment of Diabetic Foot Ulcer. Research Journal of Pharmacy and Technology. 2019;12:6 pp. 2637-2644
78. Yue Y, Wei H, Guo J, Yang Y. Ceria-based peroxidase-mimicking nanozyme with enhanced activity: A coordination chemistry strategy. Colloids and Surfaces A: Physicochemical and Engineering Aspects. 2021;610:p 125715
79. Yang K, Liu J, Yang Y, Li W. Engineering the microenvironment of Cu-MOF nanozyme via modulating ligand hydrophobicity for array-based profiling of phenolic acids in natural products. Talanta. 2026;302:p 129350
80. Fayad E, Binjawhar DN, Alqurashi M, Hadadi F, Alhomodi AF, Katouah HA, Alghanem MS, Khan KA. Biochar and Cerium Oxide Nanoparticles Mitigate Nickel and Drought Stress in Chickpea (Cicer Arietinum L.): Roles of Microbiome, Health Risk, and Molecular Responses. Journal of Soil Science and Plant Nutrition. 2026
81. Lu H, Liu Z, Wang Y, Casals E, Hanzu FA, Morales-Ruiz M, Zeng M, Casals G. Endocrine nanozymology: Nanozyme applications in diabetes, obesity, and hormonal disorders. Theranostics. 2026;16:2 pp. 712-735
82. Ernst S, Buckland J, Meredith S, Tapsak MA. Year-Long Measurement of Glucose Oxidase Deactivation in Electrochemical Glucose Sensors. IEEE Sensors Journal. 2026
83. Samukaite -BU, Mazetyte -SR, Chernyakova K, Karpicz R, Ramanavicius A. Time-resolved fluorescence spectroscopy based evaluation of stability of glucose oxidase. International Journal of Biological Macromolecules. 2020;163:pp 676-682
84. N. Verma. Risk assessment studies of the impact of occupational exposure of pharmaceutical workers on the development of antimicrobial drug resistance. Journal of Occupational Environmental Hygiene. 2020;17:no 10, pp. 437-446
85. Lin Y-H, Shen L-J, Chou T-H, Shih Y-H. Synthesis, Stability, and Cytotoxicity of Novel Cerium Oxide Nanoparticles for Biomedical Applications. Journal of Cluster Science. 2021;32:2 pp. 405-413
86. Barker E, Shepherd J, Asencio IO. The Use of Cerium Compounds as Antimicrobials for Biomedical Applications. Molecules. 2022;27:9 Art. no. 9
87. Zandi M, Hosseini F, Adli AH, Salmanzadeh S, Behboudi E, Halvaei P, Khosravi A, Abbasi S. State-of-the-art cerium nanoparticles as promising agents against human viral infections. Biomedicine & Pharmacotherapy. 2022;156:p 113868
88. Farias IAP, Dos SCCL, Sampaio FC. Antimicrobial Activity of Cerium Oxide Nanoparticles on Opportunistic Microorganisms: A Systematic Review. BioMed Research Internasional. 2018: 1, p. 1923606.
89. Putri GE, Rilda Y, Syukri S, Labanni A, Arief S. Highly antimicrobial activity of cerium oxide nanoparticles synthesized using Moringa oleifera leaf extract by a rapid green precipitation method. Journal of Materials Research and Technology. 2021;15:pp 2355-2364
90. Pertiwi MBB, Indahyani DE, Praharani D. Level Glukosa Darah pada Mencit Diabetes Setelah Pemberian Ekstrak Rumput Laut Coklat (Phaeophyta). Pustaka Kesehatan. 2021;9:2 pp. 84-89
91. S J, Babu M. Knowledge of Diabetic Clients Regarding Self-care Practices in Management of type II Diabetes Mellitus at selected Rural Community of Bangalore, Karnataka. Asian Journal of Nursing Education and Research. 2017 7, 1, pp. 86-94
Author contact
Corresponding author: Suhailah Hayaza; Email: suhailahunair.ac.id.

 Global reach, higher impact
Global reach, higher impact