Nanotheranostics 2018; 2(2):184-196. doi:10.7150/ntno.22114 This issue Cite
Research Paper
Penicillin Detection by Tobacco Mosaic Virus-Assisted Colorimetric Biosensors
1. Institute of Biomaterials and Biomolecular Systems, University of Stuttgart, 70569 Stuttgart, Germany
2. Institute of Nano- and Biotechnologies, FH Aachen, Campus Jülich, 52428 Jülich, Germany
3. Institute of Complex Systems (ICS-8), Forschungszentrum Jülich GmbH, 52525 Jülich, Germany
Received 2017-7-27; Accepted 2017-12-23; Published 2018-2-20
Abstract
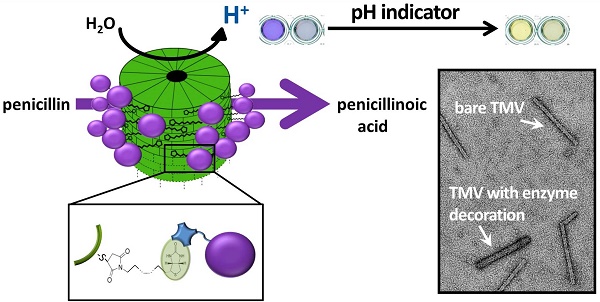
The presentation of enzymes on viral scaffolds has beneficial effects such as an increased enzyme loading and a prolonged reusability in comparison to conventional immobilization platforms. Here, we used modified tobacco mosaic virus (TMV) nanorods as enzyme carriers in penicillin G detection for the first time. Penicillinase enzymes were conjugated with streptavidin and coupled to TMV rods by use of a bifunctional biotin-linker. Penicillinase-decorated TMV particles were characterized extensively in halochromic dye-based biosensing. Acidometric analyte detection was performed with bromcresol purple as pH indicator and spectrophotometry. The TMV-assisted sensors exhibited increased enzyme loading and strongly improved reusability, and higher analysis rates compared to layouts without viral adapters. They extended the half-life of the sensors from 4 - 6 days to 5 weeks and thus allowed an at least 8-fold longer use of the sensors. Using a commercial budget-priced penicillinase preparation, a detection limit of 100 µM penicillin was obtained. Initial experiments also indicate that the system may be transferred to label-free detection layouts.
Keywords: tobacco mosaic virus (TMV), antibiotic detection, pH indicator, viral enzyme nanocarrier, penicillinase, enzyme shelf-life
Introduction
Since the discovery of penicillin G (the world's first antibiotic; a β-lactam antibiotic synonymously known as benzylpenicillin) in 1928 by Sir Alexander Fleming [1], antibiotics have revolutionized the world of modern medicine. The treatment of wounds and infections with penicillin has been saving millions of people around the world. Moreover, antibiotics are being used to a great extent in livestock production. Due to the extensive use in animal husbandry and inappropriate prescribing of antibiotics in human medicine, many types of bacteria have developed resistance; some have even become resistant to more than one type of antibiotic (reviewed e.g. in [2, 3]). The antibiotic resistance of disease-causing bacteria is of particular concern since they can be transmitted to humans both by animal products and by daily contact with other humans, or upon medical therapy. The incidence and spread of different resistance mechanisms is threatening our ability to treat even common infections, which can result in prolonged illness, epidemic/pandemic spread of infections, use of more toxic drugs and increasing mortality rates of infectious diseases [4, 5]. Therefore, it is important to support efforts to minimize the inappropriate use of antibiotics and to monitor the antibiotic residues in wastewater, offal and animal products, especially when entering the human food chain via meat, milk and eggs. In order to control this threat, the European Union (EU) has issued strict regulations including specific maximum residue limits (MRLs) for each veterinary drug in animal husbandry (Council Regulation 2377/90/EEC). For instance, the MRL of benzylpenicillin (penicillin G) in milk is 4 µg/kg (≈ 12 nM) [6, 7]. A multitude of analytical methodologies has been developed (as e.g. comprehensively reviewed in [8-13]). Some methods were designed to easily yield clinically relevant information on whether or not bacteria from hospitals can degrade antibacterial drugs and thus are resistant to those. Important detection methods are based on the cleavage of the β-lactam ring of antibiotics such as penicillin and cephalosporin by the bacterial enzyme β-lactamase (Figure 1A). The activity of β-lactamases, like penicillinase, has been assayed by a variety of chemical methods such as manometric techniques (detection of CO2 evolution in bicarbonate buffers) [14, 15], growth inhibition, and a diversity of chromogenic methods, including acidometric methods with pH indicators (halochromic dyes [16, 17]; Figure 1C), or directly with chromogenic substrates (e.g. chromogenic cephalosporin [18]; Figure 1D), or iodometric assays (compared e.g. in [18-20]; Figure 1B) in combination with spectrophotometry. Acidometric methods, as used in this work, are based on the formation of penicilloic acid with a pH change, monitored by a pH indicator in a low-buffered system (Figure 1C). The halochromic indicators mostly applied are phenol red (e.g. [17, 20-23]) and bromcresol purple ([19, 24, 25], for review [12]).
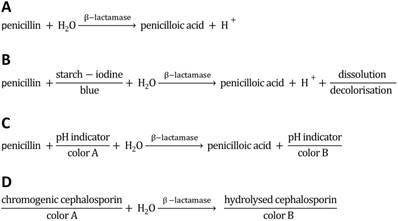
Reaction scheme of penicillinase activity and schematics of acidometric penicillin G detection (this work), and detection methods based on starch-iodine and chromogenic substrates. A) Hydrolysis of penicillin catalyzed by penicillinase. B) Reaction scheme of penicilloic acid and starch-iodine solution (iodometric assay). C) Reaction scheme of acidometric detection of penicillinase activity based on a color change of a halochromic pH indicator. D) Scheme of a spectrophotometric detection based on chromogenic substrates (e.g. chromogenic cephalosporin).
Virus particles such as tobacco mosaic virus (TMV), cowpea mosaic virus (CPMV), cowpea chlorotic mottle virus (CCMV) or the bacteriophage M13 are frequently used biotemplates for efficient analyte detection in sensing devices (as e.g. reviewed in [26-29]). These particles offer well-defined, nanostructured and stable protein backbones enabling the multivalent presentation of functional groups. Moreover, certain plant viruses and phages are attractive carrier scaffolds also due to their ease of production in high yield, and their lack of toxicity and pathogenicity to mammals. Wildtype or engineered M13 has, for instance, been used in the colorimetric detection of volatile organic compounds, in humidity sensors [30], and for the detection of explosives such as trinitrotoluene (TNT) [31] or for tracing of antibiotics [32]. It was further employed for e.g. the detection of pathogens such as Salmonella spec. or Bacillus anthracis (reviewed in [27]). Dual-functionalized hepatitis B virus (HBV) was successfully used as a signal-amplifying adapter particle promoting high sensitivity detection of troponin, a marker for acute myocardial infarction [33]. CPMV was utilized as a platform for signal enhancement in direct and sandwich immunoassays [34, 35], for the detection of staphylococcal enterotoxin B [36] or as framework facilitating avidin detection [37]. TMV was successfully applied as viral carrier in sensor systems for TNT [38], and for selective antibody recognition in both optical micro-disc resonator [39] and capillary microfluidics-integrated impedance sensor systems [40], and has also been used in a thin film sensor for the detection of volatile organic compounds [41]. In combination with inorganic compounds, TMV nanoparticles coated with palladium as sensing material have promoted the development of a surface acoustic wave (SAW) hydrogen sensor [42].
In this study, a biosensor employing TMV-based enzyme carrier scaffolds was developed for acidometric detection of penicillin. The incorporation of viral adapters was pursued, as it has been shown recently that biosensors can profit from TMV nanotubes also as enzyme immobilization supports [43, 44]. TMV is a highly stable, rod-shaped plant virus with a precisely structured protein capsid. It has a length of 300 nm, a diameter of 18 nm and an inner channel of 4 nm diameter. The capsid, with 2130 coat protein (CP) subunits, can be conjugated with functional molecules by appropriate chemistry, facilitated by CP variants with highly reactive coupling sites. TMV has acted as a well-defined high-yield nanotemplate in proof-of-concept experiments for a multitude of possible applications, reviewed in [45]. Promising results were obtained with full TMV sticks serving as nanocarriers for enzymes [43]. The two-enzyme cascade system of glucose oxidase (GOx) and horseradish peroxidase (HRP) was successfully installed on the viral adapter rods, enabling a colorimetric detection of glucose in combination with a suitable chromogenic co-substrate. A TMV mutant (TMVCys) exposing thiol groups on its surface was functionalized with bifunctional maleimide-PEG11-biotin linkers, allowing a dense decoration with active streptavidin ([SA])-conjugated enzymes. Besides enrichement of enzymes on sensor surfaces, additional beneficial effects of TMV rods were found: GOx and HRP exhibited an increased reusability, a greater storage stability and a higher regenerability [43]. Thus, a great potential application spectrum of virus-enzyme conjugates in biosensors and related bioanalytical devices is emerging. TMV enzyme carriers were also evaluated as adapters on platinum (Pt) electrode arrays on sensor chips in electrochemical biosensors for amperometric glucose detection, which effectuated a superior sensor performance [44].
The main goal, now, was to test whether the TMV nanoadapters could be functionalized with penicillinase similarly. The performance of the TMV-assisted antibiotic sensing layouts was determined, primarily in a classic colorimetric setup.
Material and Methods
If not otherwise stated, all chemicals were purchased from Roth (Karlsruhe, Germany). Concentrations of proteins and virus particles were determined by UV spectrometry (Nanodrop, Peglab, Erlangen, Germany). Graphs and diagrams were prepared by use of Inkscape (Software Freedom Conservancy, Brooklyn, NY, USA) and Graph Pad Prism 4 (Graph Pad Software Inc., San Diego, CA, USA). A list of the most common abbreviations is provided in Table 1. Note: The activity of the enzyme/enzyme-conjugates is measured in International Units (U) using penicillin G as substrate.
List of most common abbreviations.
| Abbreviation | Description |
|---|---|
| TMV | Tobacco mosaic virus |
| TMVCys | S3C-containing mutant of TMV, exposing thiol groups on every CP |
| TMVCys/Bio | TMVCys equipped with [maleimide-]-PEG11-biotin linkers (functionalized particles) |
| CP | coat protein |
| [SA] | streptavidin |
| Pen | penicillinase |
| [SA]-Pen | streptavidin-conjugated Pen |
TMV Isolation and Functionalization
TMV isolation: TMVCys particles (genetically modified to expose a thiol group on each CP [46]) were isolated from infected tobacco plants (N.tabacum 'Samsun' nn) according to Chapman [47], using 1-butanol extraction followed by PEG precipitation and ultracentrifugation (Beckman Coulter Optima L90K, 45 Ti rotor, 34 000 rpm ≈ 134,000 g (RCFmax), 4 °C, 2 h). Solutions of isolated virus (5 - 10 mg/ml) were stored in 10 mM sodium potassium phosphate buffer (SPP) pH 7.0 at 4 °C.
Particle functionalization: Bifunctional biotin linkers (EZ-Link® Maleimide-PEG11-Biotin; ThermoScientific, Rockford, IL) were aliquoted under protective atmosphere and stored at 4 °C until shortly before use. Prior to use, a small amount of linker was dissolved in dimethylsulfoxid (DMSO). For functionalization (biotinylation) of virus particles, maleimide-biotin linkers were added in 3-fold molar excess (calculated for 2130 CPs per TMV nanotube) to a TMVCys suspension and incubated at 30 °C for 3 h. Unbound linker molecules were separated from TMVCys/Bio (functionalized particles) by ultracentrifugation (Beckman Coulter Optima L90K, 45Ti rotor, 34 000 rpm ≈ 134,000 g (RCFmax), 4 °C, 2 h). Pellets containing TMVCys/Bio were resuspended in 10 mM SPP buffer pH 7.0 and further analyzed by 15 % SDS-PAGE [48]. SDS-gels, stained with Coomassie Brilliant Blue R250 (Serva Electrophoresis, Heidelberg, Germany) were used to determine the coupling efficiency of linker to CP using the image evaluation software ImageJ (Rasband, 1997-2010).
Conjugation of Streptavidin and Penicillinase
The enzyme penicillinase (Pen) from Bacillus cereus was purchased as lyophilized powder containing ~ 3 - 5 % protein with phosphate and citrate buffer salt (P0389, Sigma-Aldrich, München, Germany). It was diluted by addition of 1 x PBS to 1 mg protein/ml. A LYNX Rapid Streptavidin Antibody Conjugation Kit® (LNK162STR, Bio-Rad AbD Serotec GmbH, Puchheim, Germany) was used for rapid conjugation of streptavidin [SA] to Pen (molar ratio [SA]:Pen = 30:1), resulting in [SA]-Pen. To verify conjugation of [SA] and Pen, samples were analyzed by 12 % SDS-PAGE [48] and stained with Coomassie Brilliant Blue R250 (Serva Electrophoresis, Heidelberg, Germany).
TEM Analysis of Enzyme-Decorated TMV Nanocarriers
TMVCys/Bio-[SA]-Pen conjugates were analyzed by transmission electron microscopy (TEM). To this end, 10 µg TMVCys/Bio were incubated with [SA]-Pen in a molar ratio of [SA]-Pen:CPCys/Bio = 1:1.5 at room temperature (RT) for 16 h. Samples were mixed in a one-to-one TMV particle ratio with undecorated TMVCys/Bio. A 0.2 µg/µl dilution of this mixture was adsorbed to Formvar®-coated, carbon-sputtered 400 mesh copper grids (Ted Pella, Redding, CA) for 10 min. Grids were washed with ultrapure water (ddH2O, 18.3 MΩ cm, purified by a membraPure system, Aquintus, Bodenheim, Germany) and negatively stained using three drops of 2 % (w/v) uranyl acetate. After removal of residual staining solution with filter paper and drying, samples were imaged with a Tecnai G2 Sphera Fei TEM (Fei, Eindhoven, Netherlands) at 120 kV and images recorded by a 16 megapixel camera (TemCam-F416 [4k × 4k], TVIPS, Gauting, Germany).
Optimization of an Acidometric (Colorimetric) Detection Method
Substrate solutions: The enzyme substrate penicillin G (penG; sodium salt; P3032, Sigma-Aldrich, München, Germany) was dissolved in ultrapure water to different concentrations ranging from 10 µM to 50 mM penG and the pH adjusted to pH 8.0.
pH indicator solutions: Different pH indicator solutions were prepared and stored in the dark at RT (0.1 % (w/v) phenol red in 20 % (v/v) ethanol (EtOH); 0.1 % (w/v) bromcresol purple in 20 % (v/v) EtOH; 0.1 % (w/v) methyl red in 90 % (v/v) EtOH; 0.1 % (w/v) bromthymol blue in EtOH; 0.1 % (w/v) cresol red in 50 % (v/v) EtOH; 0.1 % (w/v) phenolphthalein in 50 % (v/v) EtOH).
Absorption spectra: To obtain absorption spectra (300 nm - 700 nm) of pH indicators, 200 µl of testing solutions (0.5 x phosphate buffered saline (PBS)) with pH values ranging from 1 to 12 were mixed with 5 µl of the particular pH indicator in microtiter plates (96-well clear polystyrene; Greiner Bio-One, Frickenhausen, Germany) and analyzed in a spectrophotometer (Spectrafluor Plus infinite M200 pro, TECAN). For each halochromic pH indicator, a specific wavelength with a maximal change in absorption over the tested pH range was chosen (phenol red: 558 nm, bromcresol purple: 588 nm, methyl red: 516 nm, bromthymol blue: 592 nm, cresol red: 572 nm, phenolphthalein: 555 nm) and used for further testing.
Comparison of pH indicators: To directly compare the suitability of the different pH indicators under Pen reaction conditions, varying enzyme amounts (5 ng ≈ 12.5 µU; 10 ng ≈ 25 µU, 15 ng ≈ 37.5 µU; 20 ng ≈ 50 µU, 25 ng ≈ 62.5 µU, all diluted in 40 µl of 0.5 x PBS) were mixed with 40 µl of 10 mM penG or 20 mM penG (final concentrations: 5 mM penG and 10 mM penG in 0.25 x PBS, respectively) and 3 µl pH indicator. The absorption of each mixture was measured at the pH indicator-specific wavelength for 15 min. The absorption change (OD/min) of each approach was inferred from the slope of the linear section. All tests were performed in repeated independent experiments each with triplicates for every penG concentration.
Detection of different β-lactam antibiotics: To evaluate the ability of Pen from B. cereus (see above) to detect distinct β-lactam antibiotics, six different antibiotics (each in 100 µl solutions, between 0 mM and 50 mM) were supplemented with 50 mU or 100 mU Pen and mixed with 3 µl bromcresol purple. The following six antibiotics were tested in direct comparison: five penicillins, i.e. penicillin G (benzylpenicillin; PubChem CID: 5904), tricarcillin (PubChem CID: 36921), carbenicillin (PubChem CID: 20824), ampicillin (PubChem CID: 6249), and cloxacillin (PubChem CID: 6098), and one cephalosporin, i.e. cefotexim (PubChem CID: 5742673). For the analyses, all antibiotics were dissolved in 0.25x PBS. The absorption changes of each reaction mixture were measured at 588 nm for 15 min.
Enzyme Coupling onto Polystyrene-Adsorbed Viral Adapters and Determination of Catalytic Activities
Adsorption of viral carriers and enzyme coupling: 5 µg biotinylated TMVCys nanorods (TMVCys/Bio) were diluted in 100 µl binding buffer (10 mM SPP pH 7.8 and 137 mM NaCl) and incubated in microtiter plates (clear F-bottom 96-well polystyrene high binding MICROLON®; No.655061, GreinerBio-One, Frickenhausen, Germany) at 4 °C for 16 h to allow adsorption of viral adapters to the well surfaces, followed by three washing steps (RT, 5 min, 1 x PBS-T (PBS, 0.05 % Tween-20)). In approaches designed to discriminate between specific (streptavidin-biotin mediated) and unspecific (mere adsorption) binding, non-specific adhesion of enzymes was reduced by blocking the wells each with 200 µl 2 % (w/v) bovine serum albumin (BSA) in 1 x PBS for 1 h at RT, followed by three consecutive washing steps as before. If not otherwise stated, these pretreated cavities were incubated with 1 U [SA]-Pen or 1 U Pen, respectively, (≈ 0.4 µg enzyme) diluted in 100 µl 1 x PBS, for 3 h at RT. After enzyme immobilization, wells were washed thoroughly 5-6 times as before.
Determination of catalytic activities: For determination of the catalytic activity in each approach, 100 µl substrate mix (10 mM penG in 0.25 x PBS pH 8.0 and 3 µl bromcresol purple) were applied. Absorption was measured for each well at 588 nm for 10 min. The absorption was plotted vs. time and the absorption change (OD/min) was derived from the slope of the linear section. All tests were carried out repeatedly with triplicates for every approach.
Reusability: After preparation of sensor wells as described above, initial catalytic activities were determined, followed by three consecutive washing steps and storage in 1x PBS at 4 °C. Remaining activities were measured for each approach every 2-3 days in repeated experiments with triplicates of each sensor well layout. Initial activities were set to 100 % and the percentages of remaining activities were calculated and plotted versus the testing period.
Detection limit: For determining the detection limit of the TMV-assisted penicillin sensors, wells were prepared as described before and treated with substrate solutions with different penG concentrations ranging from 0 mM to 50 mM (triplicates, with a few duplicates; pH adjusted to pH 8.0 before use). The absorption was measured for each well at 588 nm for 10 min and the absorbance change (OD/min) determined. Differences occurring among the the distinct analyte concentrations were subjected to statistical evaluation by unpaired t-testing. A p value of less than 0.05 was considered to indicate a significant difference between the absorbance changes reached with varying penG amounts (* p < 0.05; ** p < 0.01; *** p < 0.001).
Results
Colorimetric detection of penicillin was performed with enzme-based biosensors employing TMV-derived nanoadapter scaffolds. Streptavidin conjugation of a commercial penicillinase and its binding to biotin-linked TMV rods enabled selective high-surface density immobilization.
Comparison of pH Indicators
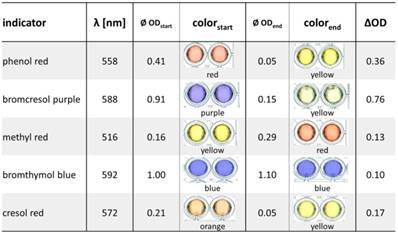
The colorimetric detection of penicillinase activity is based on acidometry employing the hydrolysis of penicillin with differential halochromic color changes of pH indicators (Figure 1C). To identify the most suitable pH indicator, six pH indicator dyes were tested: phenol red, bromcresol purple, methyl red, bromthymol blue, cresol red and phenolphthalein, which showed a clear color change between pH 3 and pH 10 (the expected working range of the penicillinase biosensor) (Figure S1). To quantify the proton generation, the absorption spectra of each pH indicator between pH 1 and pH 12 were measured to determine the wavelength of maximal change (Table 2, Figure S2). Phenolphthalein did not show any relevant color change in the analyte/enzyme mixtures. Best changes (OD/min) (Figure S3) and ∆OD values (ODstart - ODend) (Table 2) were obtained by bromcresol purple (∆OD = 0.76), followed by phenol red (∆OD = 0.36), cresol red, methyl red and bromthymol blue (∆OD values from 0.1 to 0.17). With bromcresol purple, low enzyme amounts (5 ng) in low penicillin concentrations (5 mM penG) were detectable (Figure S3).
Comparison of absorption changes of different pH indicators at the optimum wavelengths. 25 ng penicillinase blended with 10 mM penG were mixed with different pH indicator solutions. Absorption of each mixture was measured at the beginning of the enzymatic reactions (ODstart) and after t = 15 min (ODend).
Recognition of different β-lactam antibiotics
The penicillinase from B. cereus was further tested for its ability to detect different β-lactam antibiotics in the two most important β-lactam classes, penicillins and cephalosporins [49], all of which are used for medical treatments. The core structure of penicillins is a β-lactam ring condensed to a sulfur-containing five-membered heterocyclic ring (i.e. a 4,5-ring system), whereas in cephalosporins, the corresponding heterocycle is six-membered (to give a 4,6-ring system), which inhibits cleavage by most penicillinase enzymes [50]. Besides penicillin G (benzylpenicillin), a naturally occurring broad-spectrum antibiotic, five semi-synthetic antibiotics with various side chains were tested: four additional penicillins, namely tricarcillin, carbenicillin, ampicillin and the penicillinase-resistant cloxacillin [51], and the cephalosporin cefotexim (Figure S5). Tricarcillin and carbenicillin are broad-spectrum carboxybenzylpenicillin derivatives, ampicillin is an aminopenicillin derivative. Cefotaxim belongs to the third-generation broad-spectrum cephalosporins. As expected, the penicillinase employed was neither able to recognize cloxacillin due to steric hindrance of the synthetic side chains [51], nor cefotaxime due to the cephalosporin's 4,6-ring system. Carbenicillin and tricarcillin were readily detectable, though with lower sensitivity (corresponding to about 50 % of that reached with penicillin G, see Figure S5). Ampicillin detection was feasible to a limited extent upon use of of higher enzyme input (about twice the amount needed as for the other antibiotics; Figure S5, B). Taken together, the B. cereus penicillinase was able to detect three distinct, pharmaceutically applied penicillin derivatives with different efficiencies, rendering it likely that also further natural and semisynthetic β-lactam antibiotics may be accessible to the sensor system. To characterize it in more detail and to evaluate the effect of TMV adapter scaffolds on the biosensor performance, penicillin G was chosen as test target, as described in the following.
Functionalization of TMV Nanocarrier and Coupling of Penicillinase
Adsorptive binding of biotinylated TMV to the sensor surfaces of microtiter plate wells was followed by enzyme immobilization by streptavidin-biotin binding (Figure 2A). To this aim, the most surface-accessible cysteine residues of the TMVCys particles were modified with maleimide-PEG11-biotin linker molecules (TMVCys/Bio), with a biotinylation efficiency of 95 % corresponding to ≈2000 biotin anchors per virus rod (Figure S4A). To allow a strong and specific coupling of Pen, enzyme molecules were conjugated with streptavidin ([SA])-complexes, resulting in [SA]-Pen (conjugation effiency: > 90 % of the Pen molecules (Figure S4B)).
Coupling and activity of streptavidin-conjugated penicillinase ([SA]-Pen) on functionalized TMV adapters. A) Illustration of the functionalization procedure of TMVCys/Bio nanotubes with [SA]-Pen conjugates. B) TEM image of a mixed sample containing bare and [SA]-Pen-decorated TMVCys/Bio adapters, blended for TEM analysis only, in order to comparatively visualize the differences between bare and enzyme-loaded biotinylated TMV (as indicated; for activity tests, pure preparations were applied). Enzyme coupling results in an increased diameter as well as intensified staining with 2 % uranyl acetate. Scale bar: 200 nm. C) Left: Input of 60 µU penicillinase (Pen) and 60 µU [SA]-Pen activity in reaction solution (10 mM penG in 0.25 x PBS pH 8.0 and 3 µl bromcresol purple/100 µl) results in similar absorption changes over time, indicating that conjugation with [SA] does not impair the enzyme activity. Right: Approaches using immobilized TMV adapters: TMVCys/Bio particles were adsorped to polystyrene plate cavities, followed by BSA blocking and application of [SA]-Pen in (A) and Pen without [SA] in (C), respectively. Plate cavities without viral adapters were blocked with BSA, followed by application of [SA]-Pen in (B) and Pen in (D), respectively.
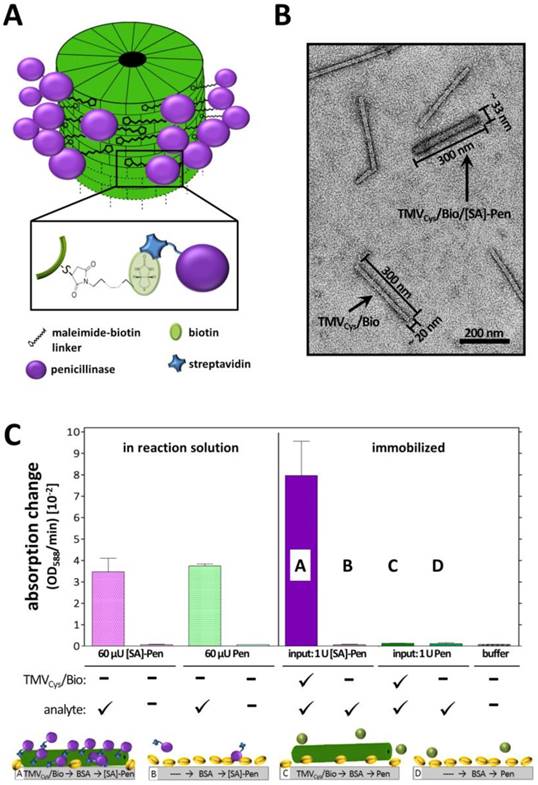
The penicillinase activity was not impaired by the streptavidin conjugation, since 60 µU of both [SA]-conjugated and free penicillinase in solution yielded equal absorption changes of ≈3 OD/min in the reaction mixture (Figure 2A). On polystyrene microtiter plates immobilized TMVCys/Bio sticks demonstrated the requirement of streptavidin for successful coupling of penicillinase. Both biotin equipment of the TMV adapters as well as streptavidin conjugation of the penicillinase was crucial for immobilization of significant amounts of enzymes, resulting in detectable substrate conversion rates (Figure 2A). The successful enzyme decoration of TMVCys/Bio sticks was verified by TEM of freely suspended particles prepared in parallel. Enzyme conjugation to TMV adapter sticks was evidenced by an increased diameter of 30 - 33 nm and intensified staining for the TMV adapter sticks (Figure 2B).
TMV-assisted Penicillin Sensor
The catalytic activities of Pen and [SA]-Pen (measured as OD/min) were analyzed after immobilization on solid supports in direct comparison between setups based on direct (adsorptive, without carrier) binding and setups equipped with biotinylated TMV nano-carriers as adapters. Equal input amounts of [SA]-Pen or Pen (each 1 U) were applied for binding into microtiter plate cavities as depicted in Figure 3: TMVCys/Bio tubes (No. 1, 5) (5 µg), TMVCys/Bio tubes (5 µg) followed by BSA treatment to reduce unspecific enzyme binding (No. 2, 6), bare plate cavities (No. 3, 7) or cavities pretreated with BSA (No. 4, 8; see also Figure 3C). Following optional TMV adapter binding, optional BSA blocking, subsequent incubation with enzyme or enzyme [SA]-conjugate, and removal of unbound [SA]-Pen or Pen thereafter, absorption changes reflecting the enzyme activity in each approach were determined by spectrophotometry. Highest absorption changes over time were obtained in TMV-assisted layouts enabling [SA]-Pen bioaffinity coupling. Cavities equipped with TMVCys/Bio (No. 1) reached a maximum change of 13 × 10-2 OD/min (Figure 3A), due to enzyme immobilization primarily through biotin-streptavidin interaction. The use of TMVCys/Bio and BSA blocking (No. 2) achieved a slightly lower absorption change of 11 x 10‑2 OD/min, indicating that in this layout, solely biotin-specific immobilization of [SA]‑Pen occurred (and thus, a minor additional amount of enzyme had adsorbed to the non-blocked polystyrene surface in layout No. 1). The blocking efficiency of BSA was confirmed in the cavities with BSA blocking preceding enzyme incubation (No. 4, 8), which did not exhibit any enzyme activity after washing. In layouts lacking either the viral adapter (No. 3, 7), or streptavidin-conjugation of the enzyme (No. 5, 6), only lower absorption changes of around 8-9 x 10-2 OD/min were obtained, underlining the efficiency of the TMV-biotin-streptavidin enzyme trapping method.
Absorption changes reflecting the catalytic activities in eight different sensor layouts and their reusability. A) Initial absorption changes (OD/min) obtained in the distinct sensor layouts upon equal input (1 U each) of [SA]-Pen or Pen, respectively. Layouts made use of TMVCys/Bio tubes (No. 1, 5), TMVCys/Bio tubes followed by BSA blocking (No. 2, 6), bare plate cavities (No. 3, 7) and BSA-blocked cavities (No. 4, 8) (schematically shown in C). B) Activities of penicillin sensors upon repeated use over five weeks. Initial activities (as in A) of each layout were set to 100 % and the remaining percentages calculated (for details refer to text). C) Cartoon of immobilization methods used for enzyme coupling in tested layouts. Same activity amounts of [SA]-Pen or Pen, respectively (each 1 U), were applied for enzyme binding to TMVCys/Bio tubes (No. 1, 5), TMVCys/Bio tubes plus BSA blocking (No. 2, 6), bare plate cavities (No. 3, 7) and cavities pretreated with BSA (No. 4, 8). Dotted line: 50 % remaining activity indicating enzyme half-life.
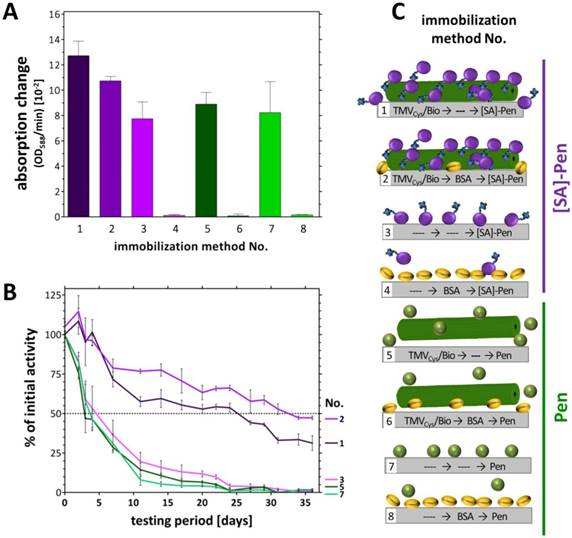
In addition to concentrating enzymes at technically relevant sites, certain immobilization routes and support materials were also shown to increase the storage stability and reusability of the bioactive molecules [52-54]. Our recent studies have revealed a remarkable stabilization of the glucose sensor bienzyme system [SA]-GOx/[SA]-HRP immobilized on biotin linker-equipped TMV nanorod scaffolds [43]. To investigate whether TMV adapters exert a similar beneficial effect also on penicillinase-conjugates coupled the same way, the layouts (No. 1‑8) were tested for their reusability every 2-3 days over a period of 5 weeks. Initial activities (Figure 3A) were set to 100 % and remaining activities after each consecutive use calculated (Figure 3B). Notably, during the whole testing period with multiple uses, the [SA]-Pen enzymes on TMVCys/Bio stick adapters showed only a small, rather continuous decrease in substrate conversion efficiency, reaching a remaining activity of 35 % up to 50 % after five weeks, best preserved in the presence of BSA (No. 1,2). This striking performance outcompeted every other layout tested. Those relying merely on non-specific adsorption (without TMV and BSA) showed a fast decrease in Pen activity with a half-life of 4‑6 days only (No. 3,7). After 3 weeks of parallel testing, less than 10 % of the initial activity, and after 5 weeks, no residual activity was detectable in the TMV-free (No. 3,7) and the TMV-supplemented but enzyme non-binding control layouts (No. 5,6). Cavities pre-treated with BSA did not adsorb substantial amounts of freely suspended enzymes at all (No. 4,8; see Figure 3A above).
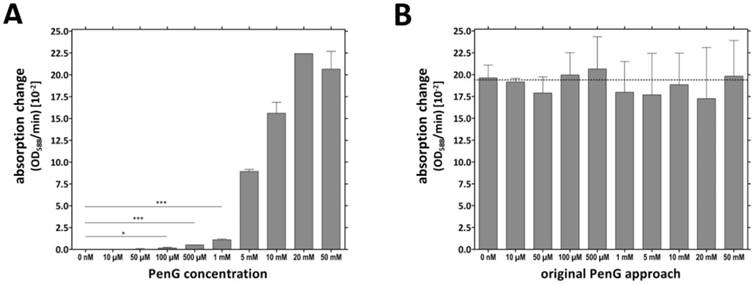
The TMVCys/Bio-[SA]-Pen system was investigated for its penicillin detection limit (limit of detection: LOD). Several microtiter plate cavities were equipped in parallel with TMVCys/Bio tubes followed by BSA blocking of unspecific enzyme binding sites (immobilization method No. 2, Figure 3C) prior to [SA]-Pen application. The TMV-assisted penicillin sensors exhibited a LOD of 100 µM penG (Figure 4A). Triplicate values for individual penG concentrations were compared by unpaired t-testing, with a p value of less than 0.05 considered to reflect a significant difference (* p < 0.05; ** p < 0.01; *** p < 0.001). This yielded distinguishable absorption changes in a detection range from 100 µM to 20 mM penG. Above this penG concentration, the system was saturated. Sensors were then washed and reused in 10 mM penG to prove a comparable functionality of all sensor wells (Figure 4B).
Penicillin detection limit reached by TMV-immobilized [SA]-Pen conjugates. A) Cavities were equipped with TMVCys/Bio tubes followed by BSA blocking (immobilization method No. 2, Figure 3C), immobilization of [SA]-Pen and spectrophotometric evaluation of the sensing performance in substrate solutions containing 0 nM - 50 mM penG and bromcresol purple. Absorption changes (OD588/min) were determined and compared to the values for substrate solutions without penicillin (0 nM) by unpaired t‑test. A p value of less than 0.05 was considered to indicate significant differences (* p < 0.05; ** p < 0.01; *** p < 0.001). Distinguishable absorption changes were obtained for ≥100 µM penG (detection limit). B) Cavities used in A were washed and re-used with a 10 mM penG solution. Similar OD/min changes were obtained for all wells, proving the comparable functionality of all sensor wells.
Electrochemical TMV-Assisted Penicillin Sensor
To put the study on a firm footing, initial tests on the performance of TMV-assisted label-free sensor layouts were conducted by means of a field-effect device (FED) (to be published in detail elsewhere). A capacitive field-effect electrolyte-insulator-semiconductor (EIS) structure was equipped with TMV adapter sticks and [SA]-Pen conjugates in an analogous manner, to enable label-free and real-time electrochemical penicillin detection in combination with TMV enzyme nanocarriers for the first time. Label-free detection setups would allow analyte detection also in turbid samples not suitable for colorimetric detection methods such as many food and environmental samples.
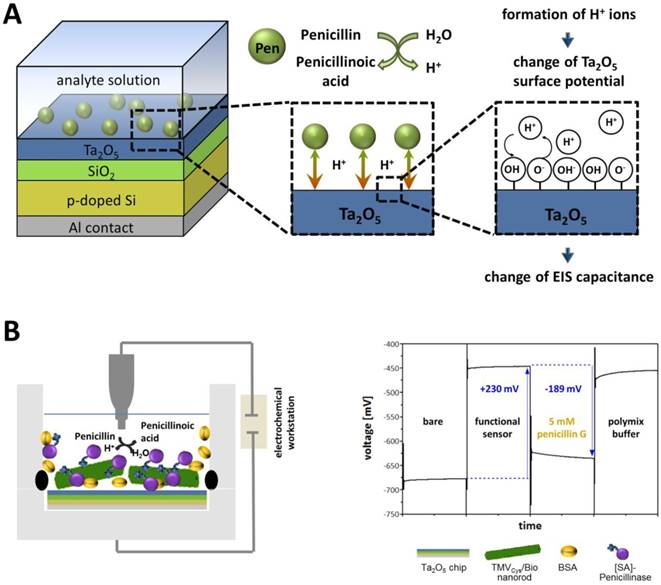
The EIS structure used for this setup represented a FED, which monitors changes in the electrical surface charge due to its pH- and ion-sensitive properties (comprehensively reviewed e.g. in [55]). The operation principle of a typical FED penicillin sensor (as e.g. used in [56-59]) is as follows: The immobilized enzymes (penicillinase) catalyze the hydrolysis of penicillin, resulting in an increasing amount of H+-ions (reaction see also Figure 1A) accruing near the sensor surface. This local pH decrease is detected by the pH-sensitive Ta2O5 surface layer (gate insulator) of the sensor chip. A resulting variation in the electrical surface charge (protonation of surface OH groups) leads to an alteration of the capacitance-voltage (C-V) and constant-capacitance (ConCap) response (see Figure 5A) of the EIS structure. The pH value decreasing due to enzymatic penG conversion and the measurable changes in the sensor signal are directly linked and thus allow a quantification of penG concentration in the analyte solution (Figure 5B).
Schematic structure and operation principle of a label-free penicillin sensor used for TMV-assisted substrate detection. A) The sensor is based on a pH-sensitive field-effect structure with Ta2O5 transducer layer. Penicillin hydrolysis achieved by penicillinase activity results in the formation of free H+ ions, which alter the Ta2O5 surface potential yielding a change of the EIS capacitance (EIS = electrolyte-insulator-semiconductor). Figure modified from [58]. B) Left: General setup for characterization of sensor performance in penicillin G-containing buffer solutions (5 mM penG). Right: Typical ConCap response curve of the developed TMV-based penicillin biosensor.
Functional TMV-assisted penicillin sensors were prepared in a stepwise manner, each step followed by electrochemical characterization by means of C-V, ConCap and impedance-spectroscopy (IS) methods. First proof-of-principle setups hind at a good sensor performance of these label-free TMV-assisted penicillin sensors (a typical ConCap response is shown in Figure 5B; further data will be published elsewhere).
Discussion
The extensive worldwide use of antibiotics in human medicine and treatment of food-producing animals results in an alarming spread of antimicrobial resistance. To overcome this unfortunate consequence, it is of utmost importance to monitor antibiotic residues in water, food and waste. Most antibiotics screening methods used nowadays depend on appropriately equipped laboratories; they are based on enzyme-linked immunosorbent assays (ELISA), a combination of liquid chromatography (LC) and mass spectrometry (MS) [60, 61], microbiological inhibition assays [62, 63] or biosensors (see e.g. reviews as [8, 11-13] and references therein).
Many of these sensors are label-free and often based on surface plasmon resonance (SPR) (see for original research e.g. [64, 65] or for review e.g. [10, 66]), quartz crystal microbalance (QCM) (reviewed in e.g. [10]), or several electrochemical techniques (potentiometric, amperometric and conductometric) [8, 10-13]. These biosensors allow high throughput, short analysis times, automation and multiple repeated uses due to high regenerability. The frequently utilized integration of biological recognition elements such as enzymes entails a high specificity and great signal amplification. Enzyme-based biosensors have achieved LODs of 1.6 µg/kg - 1.3 mg/kg (≈ 5 nM - 4 µM) as reviewed e.g. in [12]. However, the biosensor design typically suffers from two main types of drawback: the first is the instability of the biological sensing component (enzyme or antibody), which is susceptible to temperature, pH and other environmental stresses and may therefore quickly lose its activity. The second is the elaborate sensor format with respect to the size of the transducer unit, the often complex and costly signal processing/readout setup, and the corresponding personnel training needed. Hence, long-term stable biosensor units accessible to both official calibration and miniaturization due to electronic readout might constitute an important step towards future on-site monitoring for antibiotic residues.
To our knowledge, this study is the first approach to use viral scaffolds as enzyme-carriers in antibiotic biosensing, in order to potentially abate at least limitations due to biomolecule degradation. The integration of viral adapters into glucose sensors had revealed a remarkable stabilizing effect of TMV nanocarrier rods on both GOx and HRP, extending the reusability and overall stability of the sensors over weeks, and enabling a high-density immobilization of active enzymes on sensor surfaces [43, 44]. Here, we sought to find out if TMV-assisted enzyme-based penicillin sensors (both colorimetric and potentiometric) would exhibit similarly improved characteristics. The application of TMV-derived nano-adapters mediated increased enzyme loading compared to TMV-free layouts (factor: ≈ 1.6 x in microtiter plates (Figures 3)). The two-step coupling procedure of [SA]-conjugated enzymes to biotin-linkers was again suitable to allow strong and specific immobilization of active enzymes (Figure 2A). The method made use of a commercial bulk penicillinase preparation that was efficiently conjugated to streptavidin by a simple coupling reagent and retained its full activity. Dense enzyme coverage of the TMV rods was confirmed by TEM (Figure 2B). As our study aimed at a first basic evaluation whether or not enzymatic penG detection could be enhanced by the viral scaffolds, a standard reasonably-priced penicillinase was employed, not selected for maximum sensitivity in these sets of experiments, yet. Nevertheless, it allowed detecting at least three distinct penicillin derivatives, all being frequently used antibacterial drugs.
To our knowledge, this is the first work comparing six different pH indicators in the same enzyme-based sensor system, including the two most frequently used halochromic dyes for acidometric penicillin detection: phenol red and bromcresol purple. In our setup, bromcresol purple turned out to be best suited since it provoked the highest pH-dependent absorption changes in a reasonable detection range (Figure S3; Table 2). The acidometric characterization of our first-generation TMV-assisted penicillin sensors with an input of 1 U [SA]-Pen showed a LOD of 100 µM and a working range up to 20 mM penG (Figure 4). This LOD is far beyond (factor: ≈ 8000 x) the regulation maxima (≈ 12 nM). However, a commercial enzyme, not particularly selected for maximum sensitivity, was used for first proof-of-principle experiments shown in this study. The use of a high-performance enzyme with better specifity, higher activity and further optimization of measuring conditions (such as pH, buffer, temperature and input amounts) could lead to a lowering of the detection limit obtained by these sensors.
Integration of modified TMV adapter sticks achieved increased reusability, better stability, higher regenerability and higher analysis rates (Figure 3). The use of TMV enzyme carriers extended the half-life of the acidometric sensors from 4‑6 days to 5 weeks (Figure 3B). Both the GOx/HRP system tested on TMV carriers before [43] and the penicillinase applied here exhibited a similar half-life of about four days under TMV-free control conditions. However, compared to the GOx/HRP half-life prolonged about threefold (to nearly two weeks) on TMV nanocarriers, that of [SA]-Pen was increased to about five weeks, i.e. more than eightfold as obvious from Figure 3B. This impressive stabilization and preservation of the bioactive sensor elements might allow the applications of the sensors even over several months. Furthermore, a capacitive field-effect EIS structure equipped with TMV adapters, in order to test a different type of readout, allowed label-free analyte detection in penicillin-spiked samples.
Enzyme activity is often affected by the molecules' binding to solid supports. A carefully considered enzyme immobilization strategy compatible with the sensor surface of choice is therefore essential for generating biosensors with good and reliable performance and high reusability. Depending on the read-out system, biological, charge-sensitive or conductive compounds serving as adapter particles or films may improve the immobilization efficiency, analyte sensitivity or stability of the biological sensing molecule. A variety of organic and inorganic materials with different beneficial properties are frequently used for enzyme immobilization, alone or in composite materials. Important examples comprise gold nanoparticles (AuNPs), magnetic nanoparticles, graphene, and carbon nanotubes (including graphene oxide (GO), carbon nanotubes (CNT), both single-walled (SWNT) and multiwalled nanotubes (MWNT)) [67-69]. Nanocrystalline quantum dots (QDs) are among the most versatile adapter particles as they are luminescent as well as semiconducting, thus allowing different read-out processes from electronic up to optical sensing [70]. However, natural (e.g. gelatine, starch, cellulose, chitin, collagen) or synthetic polymers (including stimulus-responsive 'smart' and conductive polymers), as well as inorganic materials such as glass, quartz or silica are equally widespread, as they enable well-controlled installation routines of sensor molecules at practically relevant surface-densities [71, 72].
For sensor enzymes, a large variety of immobilization strategies is pursued for different detection layouts, which still demands for optimization and modification in every new enzyme-analyte combination and sensor system created. Recently, a cost-effective colorimetric method for the detection of cholesterol in aqueous solutions based on enzyme-modified gold nanoparticles was developed [73]. Cholesterol oxidase immobilized on AuNPs was used to produce H2O2 in proportion to the level of cholesterol being detected in solution. Due to the electron-accepting properties of H2O2, increasing analyte levels led to changes in the optical properties of the AuNPs. Successful detection of urea was achieved by help of three-dimensionally expanded template structures: layer-by-layer (LbL) assemblies of polyethyleneimine (PEI) and urease were deposited onto reduced GO (rGO) [74]. The integration of the weak polyelectrolyte PEI improved the pH response of the underlying graphene-based FET due to its transducing properties, thereby enabling a LOD of 1 μM urea. To improve the water dispersibility of rGO and to allow efficient glucose detection, rGO particles were also integrated into a nanocomposite matrix consisting of chitosan and AuNPs [75]. The composite provided a beneficial microenvironment for the immobilization of GOx on a glassy carbon electrode.
Penicillin biosensing has been achieved by a number of different sensor layouts before. An electrochemical immunosensor employed anti-penicillin G antibodies integrated in a supported bilayer lipid membrane (s-BLM) modified with AuNPs [76]. Binding of penicillin G was detected by impedance spectroscopy, with the AuNPs serving mainly as antibody-stabilizing additives. The approach was unprecedentedly sensitive, with a LOD below one femtoM, and about 85 % of the initial response still possible after one week of wet storage. Our detection layout was, as in most other previous studies, based on enzymatic recognition and conversion of the antibiotic. The penicillinase enzyme was exposed on plant virus-based nanorod adapters, which combine different advantageous properties: They increase the surface area available for sensor molecule coupling in the detector, their soft-matter surface can exert strong long-term stabilizing effects on the recognition elements' enzymatic activity, and their repetitive, multivalent protein coat with thousands of addressable sites can be used for various coupling approaches. Covalent, chemically and spatially selective linkage in combination with bioaffinity binding was employed with highly promising results in this work. In other studies, penicillinase was immobilized e.g. on field-effect sensors functionalized with layer-by-layer fabricated films containing SWNTs and polyamidoamine (PAMAM) dendrimers [56]. The integration of nanostructured, three-dimensionally applied PAMAM/SWNT LbL films resulted in penicillin biosensing covering the range from 5.0 µM to 25 mM penG. Compared to SWCNTs, single-graphene nanosheets (SGNs) exhibit a significantly higher conductivity and offer a larger surface area. Immobilization of penicillinase on electrodes with LbL films containing SGNs preadsorbed with hematein (SGN-hematein) and ionic liquids allowed high sensitivity detection of penicillin in an amperometric setup [77]. This electrochemical system was also applicable to milk samples. A similar biosensor was developed by LBL co-immobilization of MWCNTs, hematein, and penicillinase on a glassy carbon electrode [78]. It allowed the detection of 50 nM penicillin V, a LOD more than two orders of magnitude lower as reached by most conventional pH change-based biosensors. Recently, penicillinase was also combined with carbon paste for the determination of penG, allowing high sensitivity amperometric detection down to about 80 nM [79].
Our layout has not yet been developed into three-dimensional sensor materials and not been tested with high-performance enzymes so far. Nevertheless, the efficient enzyme docking, the strong stabilizing effect of the virus microenviroment on the penicillinase activity, and the resulting improvement of sensor performance in direct comparison to TMV-free layouts indicate that the plant-viral particles may become a particularly suitable and broadly applicable platform for the presentation of bioactive molecules. So far, the TMV-based enzyme carriers have now proved themselves worthwhile in two different model sensing systems (for glucose and penicillin), and in four different sensor setups. Therefore, the integration of TMV in further biosensors and lab-on-a-chip devices with different setups and other analyte targets is worth continuative testing. In the future, a toolbox of distinct viral adapter/enzyme-complexes may be created, based on variable coupling methods and biomolecular partners enabling substrate detection in numerous kinds of sensor layouts. The enrichment and stabilization of enzymes on sensor surfaces by help of the richly available, plant-produced TMV-based nanocarriers, in combination with more elaborated measuring-setups or more sensitive chip-materials, might enable a miniaturization of biosensor units to small hand-held devices in the future. This might allow month-long detection in less developed and newly industrialized countries without the need for fully equipped laboratories and special training for operators.
Conclusion
The study has demonstrated the general functionality of viral enzyme-nanocarriers in penicillin detection. TMV nanotube adapters densely equipped with penicillinase enzymes were integrated into setups allowing acidometric analyte detection. The most important results are: (1) Compared to mere adsorption, the presence of the TMV-based enzyme carriers increased selective enzyme loading of the sensor surfaces with equal enzyme input by a factor of ≈ 1.6 x. (2) TMV-assisted sensors exhibited superior performance along with a significantly increased half-life (5 weeks instead of several days), thus allowing an at least 8-fold longer use of the sensors. (3) Using a commercially available enzyme preparation, not particularly selected for maximum sensitivity, a LOD of 100 µM penG was obtained.
The integration of TMV/enzyme complexes in biosensors was shown to achieve superior working characteristics. In the future, this could be used to promote detection of a wide range of chemical and biological species in miniaturized detection devices.
Acknowledgements
The authors thank Melanie Jablonski and Thomas Bronder for their comprehensive support and for providing the technical equipment needed for electrochemical analysis. Furthermore, CK and CW thank Diether Gotthardt for taking great care of the tobacco plants, and Rebecca Hummel for assistance with the TEM. Special thanks go to Prof. Dr. Holger Jeske for intense debates and constructive support.
Supplementary Material
Supplementary figures.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Fleming A. On the antibacterial action of cultures of a penicillium, with special reference to their use in the isolation of B. influenzae. Br J Exp Pathol. 1929;10:226-36
2. Dever LA, Dermody TS. Mechanisms of bacterial resistance to antibiotics. Arch Intern Med. 1991;151:886-95
3. Blair JM, Webber MA, Baylay AJ, Ogbolu DO, Piddock LJ. Molecular mechanisms of antibiotic resistance. Nat Rev Microbiol. 2015;13:42-51
4. World Health Organization. WHO's first global report on antibiotic resistance reveals serious, worldwide threat to public health. Antimicrobial resistance-global surveillance report Virtual Press Conference. 2014
5. Ventola CL. The antibiotic resistance crisis: part 1: causes and threats. Pharm Ther. 2015;40:277-83
6. McEvoy J. Contamination of animal feedingstuffs as a cause of residues in food: a review of regulatory aspects, incidence and control. Anal Chim Acta. 2002;473:3-26
7. Lee M, Lee H, Ryu P. Public health risks: Chemical and antibiotic residues-review. Asian-Australas J Anim Sci. 2001;14:402-13
8. Kantiani L, Farré M, Barceló D. Analytical methodologies for the detection of β-lactam antibiotics in milk and feed samples. Trends Anal Chem. 2009;28:729-44
9. Toldrá F, Reig M. Methods for rapid detection of chemical and veterinary drug residues in animal foods. Trends Food Sci Technol. 2006;17:482-9
10. Ricci F, Volpe G, Micheli L, Palleschi G. A review on novel developments and applications of immunosensors in food analysis. Anal Chim Acta. 2007;605:111-29
11. Blasco C, Picó Y, Torres CM. Progress in analysis of residual antibacterials in food. Trends Anal Chem. 2007;26:895-913
12. Babington R, Matas S, Marco M-P, Galve R. Current bioanalytical methods for detection of penicillins. Anal Bioanal Chem. 2012;403:1549-66
13. Cháfer-Pericás C, Maquieira A, Puchades R. Fast screening methods to detect antibiotic residues in food samples. Trends Anal Chem. 2010;29:1038-49
14. Henry RJ, Housewright RD. Studies on penicillinase. 2. Manometric method of assaying penicillinase and penicillin, kinetics of the penicillin-penicillinase reaction, and the effects of inhibitors on penicillinase. J Biol Chem. 1947;167:559-71
15. Citri N. Two antigenically different states of active penicillinase. Biochim Biophys Acta. 1958;27:277-81
16. Samuni A. A direct spectrophotometric assay and determination of Michaelis constants for the β-lactamase reaction. Anal Biochem. 1975;63:17-26
17. Thornsberry C, Kirven LA. Ampicillin resistance in haemophilus influenzae as determined by a rapid test for beta-lactamase production. Antimicrob Agents Chemother. 1974;6:653-4
18. Sng EH, Yeo KL, Rajan VS, Lim AL. Comparison of methods for the detection of penicillinase-producing Neisseria-Gonorrhoeae. Br J Vener Dis. 1980;56:311-3
19. Lucas T. An evaluation of 12 methods for the demonstration of penicillinase. J Clinic Path. 1979;32:1061-5
20. Skinner A, Wise R. A comparison of three rapid methods for the detection of beta-lactamase activity in Haemophilus influenzae. J Clinic Pathol. 1977;30:1030-2
21. Bidya S, Suman R. Comparative Study of Three β Lactamase Test Methods in Staphylococcus aureus Isolated from Two Nepalese Hospitals. Open J Clin Diagn. 2014;4:47-53
22. Saz AK, Jackson LJ, Lowery DL. Staphylococcal Penicillinase.1. Inhibition and Stimulation of Activity. J Bacteriol. 1961;82:298-304
23. Nordmann P, Dortet L, Poirel L. Rapid detection of extended-spectrum-β-lactamase-producing Enterobacteriaceae. J Clinic Microb. 2012;50:3016-22
24. Sng E, Yeo K, Rajan V. Simple method for detecting penicillinase-producing Neisseria gonorrhoeae and Staphylococcus aureus. Br J Vener Dis. 1981;57:141-2
25. Collins J, Mason D, Perkins W. A microphotometric method for the estimation of penicillinase in single bacteria. Microbiology. 1964;34:353-62
26. Capek I. Viral nanoparticles, noble metal decorated viruses and their nanoconjugates. Adv Colloid Interface Sci. 2015;222:119-34
27. Mao C, Liu A, Cao B. Virus-based chemical and biological sensing. Angew Chem Int Ed Engl. 2009;48:6790-810
28. Soto CM, Ratna BR. Virus hybrids as nanomaterials for biotechnology. Curr Opin Biotechnol. 2010;21:426-38
29. Raeeszadeh-Sarmazdeh M, Hartzell E, Price JV, Chen W. Protein nanoparticles as multifunctional biocatalysts and health assessment sensors. Curr Opin Chem Engin. 2016;13:109-18
30. Oh J-W, Chung W-J, Heo K, Jin H-E, Lee BY, Wang E. et al. Biomimetic virus-based colourimetric sensors. Nat Commun. 2014;5:3043
31. Jin H, Won N, Ahn B, Kwag J, Heo K, Oh J-W. et al. Quantum dot-engineered M13 virus layer-by-layer composite films for highly selective and sensitive turn-on TNT sensors. Chem Commun. 2013;49:6045-7
32. Moon J-S, Park M, Kim W-G, Kim C, Hwang J, Seol D. et al. M-13 bacteriophage based structural color sensor for detecting antibiotics. Sens Actuators B. 2017;240:757-62
33. Park J-S, Cho MK, Lee EJ, Ahn K-Y, Lee KE, Jung JH. et al. A highly sensitive and selective diagnostic assay based on virus nanoparticles. Nat Nanotechnol. 2009;4:259-64
34. Sapsford KE, Soto CM, Blum AS, Chatterji A, Lin T, Johnson JE. et al. A cowpea mosaic virus nanoscaffold for multiplexed antibody conjugation: application as an immunoassay tracer. Biosens Bioelectron. 2006;21:1668-73
35. Soto CM, Blaney KM, Dar M, Khan M, Lin B, Malanoski AP. et al. Cowpea mosaic virus nanoscaffold as signal enhancement for DNA microarrays. Biosens Bioelectron. 2009;25:48-54
36. Soto CM, Martin BD, Sapsford KE, Blum AS, Ratna BR. Toward single molecule detection of staphylococcal enterotoxin B: mobile sandwich immunoassay on gliding microtubules. Anal Chem. 2008;80:5433-40
37. Blum AS, Soto CM, Sapsford KE, Wilson CD, Moore MH, Ratna BR. Molecular electronics based nanosensors on a viral scaffold. Biosens Bioelectron. 2011;26:2852-7
38. Zang F, Gerasopoulos K, Fan XZ, Brown AD, Culver JN, Ghodssi R. An electrochemical sensor for selective TNT sensing based on Tobacco mosaic virus-like particle binding agents. Chem Commun. 2014;50:12977-80
39. Fan XZ, Naves L, Siwak NP, Brown A, Culver J, Ghodssi R. Integration of genetically modified virus-like-particles with an optical resonator for selective bio-detection. Nanotechnology. 2015;26:205501
40. Zang F, Gerasopoulos K, Brown AD, Culver JN, Ghodssi R. Capillary microfluidics-assembled virus-like particle bionanoreceptor interfaces for label-free biosensing. ACS Appl Mater Interfaces. 2017;9:8471-9
41. Bruckman MA, Liu J, Koley G, Li Y, Benicewicz B, Niu Z. et al. Tobacco mosaic virus based thin film sensor for detection of volatile organic compounds. J Mater Chem. 2010;20:5715-9
42. Srinivasan K, Cular S, Bhethanabotla VR, Lee SY, Harris MT, Culver JN. AIChE Annual Meeting. Cincinnati (USA). 2005:74d /1
43. Koch C, Wabbel K, Eber FJ, Krolla-Sidenstein P, Azucena C, Gliemann H. et al. Modified TMV particles as beneficial scaffolds to present sensor enzymes. Front Plant Sci. 2015;6:1137
44. Bäcker M, Koch C, Eiben S, Geiger F, Eber F, Gliemann H. et al. Tobacco mosaic virus as enzyme nanocarrier for electrochemical biosensors. Sens Actuator B Chem. 2017;238:716-22
45. Koch C, Eber FJ, Azucena C, Forste A, Walheim S, Schimmel T. et al. Novel roles for well-known players: from tobacco mosaic virus pests to enzymatically active assemblies. Beilstein J Nanotechnol. 2016;7:613-29
46. Geiger FC, Eber FJ, Eiben S, Mueller A, Jeske H, Spatz JP. et al. TMV nanorods with programmed longitudinal domains of differently addressable coat proteins. Nanoscale. 2013;5:3808-16
47. Chapman SN. Tobamovirus isolation and RNA extraction. Methods Mol Biol. 1998;81:123-9
48. Laemmli UK. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970;227:680-5
49. Fischbach MA, Walsh CT. Antibiotics for emerging pathogens. Science. 2009;325:1089-93
50. Singh SB, Young K, Silver LL. What is an “ideal” antibiotic? Discovery challenges and path forward. Biochem Pharmacol. 2017
51. Pop E, Wu WM, Bodor N. Chemical delivery systems for some penicillinase-resistant semisynthetic penicillins. J Med Chem. 1989;32:1789-95
52. Katchalski-Katzir E. Immobilized enzymes-learning from past successes and failures. Trends Biotechnol. 1993;11:471-8
53. Mateo C, Palomo JM, Fernandez-Lorente G, Guisan JM, Fernandez-Lafuente R. Improvement of enzyme activity, stability and selectivity via immobilization techniques. Enzyme Microb Technol. 2007;40:1451-63
54. Singh RK, Tiwari MK, Singh R, Lee JK. From protein engineering to immobilization: promising strategies for the upgrade of industrial enzymes. Int J Mol Sci. 2013;14:1232-77
55. Schöning MJ, Poghossian A. Recent advances in biologically sensitive field-effect transistors (BioFETs). Analyst. 2002;127:1137-51
56. Siqueira JR Jr, Abouzar MH, Poghossian A, Zucolotto V, Oliveira ON Jr, Schöning MJ. Penicillin biosensor based on a capacitive field-effect structure functionalized with a dendrimer/carbon nanotube multilayer. Biosens Bioelectron. 2009;25:497-501
57. Abouzar MH, Poghossian A, Razavi A, Besmehn A, Bijnens N, Williams OA. et al. Penicillin detection with nanocrystalline-diamond field-effect sensor. Phys Status Solidi A. 2008;205:2141-5
58. Poghossian A, Schöning MJ, Schroth P, Simonis A, Lüth H. An ISFET-based penicillin sensor with high sensitivity, low detection limit and long lifetime. Sens Actuator B Chem. 2001;76:519-26
59. Poghossian A, Abouzar MH, Razavi A, Bäcker M, Bijnens N, Williams OA. et al. Nanocrystalline-diamond thin films with high pH and penicillin sensitivity prepared on a capacitive Si-SiO2 structure. Electrochim Acta. 2009;54:5981-5
60. Liu Z, Lu S, Zhao C, Ding K, Cao Z, Zhan J. et al. Preparation of anti-danofloxacin antibody and development of an indirect competitive enzyme-linked immunosorbent assay for detection of danofloxacin residue in chicken liver. J Sci Food Agric. 2009;89:1115-21
61. Meng M, Xi R. Review: current development of immunoassay for analyzing veterinary drug residue in foods and food products. Anal Lett. 2011;44:2543-58
62. Pikkemaat MG. Microbial screening methods for detection of antibiotic residues in slaughter animals. Anal Bioanal Chem. 2009;395:893-905
63. Pikkemaat MG, Rapallini ML, Zuidema T, Elferink JA, Oostra-Van Dijk S, Driessen-Van Lankveld WD. Screening methods for the detection of antibiotic residues in slaughter animals: comparison of the European Union Four-Plate Test, the Nouws Antibiotic Test and the Premi® Test (applied to muscle and kidney). Food Addit Contam. 2011;28:26-34
64. Gustavsson E, Degelaen J, Bjurling P, Sternesjö Å. Determination of β-lactams in milk using a surface plasmon resonance-based biosensor. J Agr Food Chem. 2004;52:2791-6
65. Gustavsson E, Bjurling P, Degelaen J, Sternesjö Å. Analysis of β-lactam antibiotics using a microbial receptor protein-based biosensor assay. Food Agric Immunol. 2002;14:121-31
66. Petz M. Recent applications of surface plasmon resonance biosensors for analyzing residues and contaminants in food. Monatsh Chem. 2009;140:953-64
67. Putzbach W, Ronkainen NJ. Immobilization techniques in the fabrication of nanomaterial-based electrochemical biosensors: A review. Sensors. 2013;13:4811-40
68. Othman A, Karimi A, Andreescu S. Functional nanostructures for enzyme based biosensors: properties, fabrication and applications. J Mater Chem B. 2016;4:7178-203
69. Ansari SA, Husain Q. Potential applications of enzymes immobilized on/in nano materials: A review. Biotechnol Adv. 2012;30:512-23
70. Holzinger M, Le Goff A, Cosnier S. Nanomaterials for biosensing applications: a review. Front Chem. 2014;2:63 -
71. Homaei AA, Sariri R, Vianello F, Stevanato R. Enzyme immobilization: an update. J Chem Bio. 2013;6:185-205
72. Datta S, Christena LR, Rajaram YRS. Enzyme immobilization: an overview on techniques and support materials. 3 Biotech. 2013:3 1-9
73. Nirala NR, Saxena PS, Srivastava A. Colorimetric detection of cholesterol based on enzyme modified gold nanoparticles. Spectrochim Acta A. 2018;190:506-12
74. Piccinini E, Bliem C, Reiner-Rozman C, Battaglini F, Azzaroni O, Knoll W. Enzyme-polyelectrolyte multilayer assemblies on reduced graphene oxide field-effect transistors for biosensing applications. Biosens Bioelectron. 2017;92:661-7
75. Ye Y, Ding S, Ye Y, Xu H, Cao X, Liu S. et al. Enzyme-based sensing of glucose using a glassy carbon electrode modified with a one-pot synthesized nanocomposite consisting of chitosan, reduced graphene oxide and gold nanoparticles. Microchim Acta. 2015;182:1783-9
76. Li H, Xu B, Wang D, Zhou Y, Zhang H, Xia W. et al. Immunosensor for trace penicillin G detection in milk based on supported bilayer lipid membrane modified with gold nanoparticles. J Biotechnol. 2015;203:97-103
77. Wu Y, Tang L, Huang L, Han Z, Wang J, Pan H. A low detection limit penicillin biosensor based on single graphene nanosheets preadsorbed with hematein/ionic liquids/penicillinase. Mater Sci Eng C. 2014;39:92-9
78. Chen B, Ma M, Su X. An amperometric penicillin biosensor with enhanced sensitivity based on co-immobilization of carbon nanotubes, hematein, and β-lactamase on glassy carbon electrode. Anal Chim Acta. 2010;674:89-95
79. do Prado TM, Foguel MV, Gonçalves LM, Maria del Pilar TS. β-Lactamase-based biosensor for the electrochemical determination of benzylpenicillin in milk. Sens Actuator B Chem. 2015;210:254-8
Author contact
Corresponding author: Christina Wege, Department of Molecular Biology and Plant Virology, Institute of Biomaterials and Biomolecular Systems, University of Stuttgart, 70569 Stuttgart, Germany, email: christina.wegeuni-stuttgart.de

 Global reach, higher impact
Global reach, higher impact