Nanotheranostics 2017; 1(1):131-140. doi:10.7150/ntno.18989 This issue Cite
Research Paper
Engineering Quantum Dots with Different Emission Wavelengths and Specific Fluorescence Lifetimes for Spectrally and Temporally Multiplexed Imaging of Cells
1. School of Electrical and Electronic Engineering, Nanyang Technological University, 639798, Singapore;
2. Changchun University of Science and Technology, Changchun, 130022, P. R. China;
3. Key Laboratory of Optoelectronics Devices and Systems of Ministry of Education /Guangdong Province, College of Optoelectronic Engineering, Shenzhen University, Shenzhen, 518060, P. R. China.
Received 2016-12-30; Accepted 2017-2-17; Published 2017-3-3
Abstract

In this work, a proof-of-concept study was performed to examine the potential of spectrally and temporally multiplexed imaging of cells by using quantum dots (QDs). The CdSe and ZAIS QDs with different emission wavelengths and well-separated fluorescence lifetimes were prepared to provide 2-dimensional information. After incubation with cells, the same type of QDs with different emission wavelengths were distinguishable in spectral imaging while different types of QDs with similar emission wavelengths but well-separated fluorescence lifetimes were resolvable in fluorescence lifetime imaging. For cells co-stained with dye and different types of QDs, the fluorescence lifetime imaging microscopy (FLIM) images showed spatially separated patterns that can be split into channel images by using the software-based time gates. Overall, the results demonstrate the feasibility of combining the 2-dimensional encoded QDs for spectrally and temporally multiplexed imaging. This method can be extended to other QDs and organic dyes to maximize the number of measurable species in multiplexed imaging and sensing applications.
Keywords: Quantum Dots, Fluorescence Lifetime, Multiplexed Imaging, Temporal Multiplexing.
Introduction
Multiplexed imaging provides a useful technique for simultaneously identifying multiple biological targets and comprehensively analyzing physiological processes at both the cellular and tissue level.[1, 2] Commonly, the fluorescent probes facilitate the acquisition of high-resolution biomedical images such as infected organs and tumor cells.[3-6] The fluorescent signals of these probes can be resolved spectrally based on their different emission wavelengths or temporally based on their specific fluorescence lifetimes, which make them applicable in spectrally or temporally multiplexed imaging. Although the spectrally multiplexed imaging has been well developed, the temporally multiplexed imaging may serve as a useful complementary method. It can be used to resolve the fluorescent probes with similar spectral wavelengths but widely differing fluorescence lifetimes, which is difficult to be discriminated in spectrally multiplexed imaging. [7, 8] By applying appropriate time gates, the image contrast can be increased due to the suppression of the scattered light and cell autofluorescence.[9-11] The combination of temporally and spectrally multiplexed imaging may increase or even multiply the number of the channels that can be resolved, thereby allowing the simultaneous detection of maximum measurable species.
In recent years, some researchers have developed fluorescent probes with resolvable fluorescence lifetimes for temporally multiplexed imaging. Hoffmann et al. encapsulated near-infrared dyes in polymer nanoparticles and obtained fluorescent probes with largely overlapped absorption/emission profiles but different fluorescence lifetimes (~1.0-3.0 ns).[7] The signals of these probes were successfully discriminated in living cells by fluorescence lifetime imaging microscopy (FLIM). For temporal multiplexing in microsecond region, Lu et al. developed a series of lanthanide encoded microspheres with tunable fluorescence lifetimes (~188-359 μs).[12] After optimizing the fitting algorithm and detection configuration suitable for the microsecond-scale lifetimes, they successfully applied the lanthanide-doped microspheres for temporally multiplexed probing of DNA strands. Quantum dots (QDs), with emissions covering UV to near-infrared region and fluorescence lifetimes ranging from nanosecond to millisecond scale,[13, 14] are promising candidates for spectrally and temporally multiplexed imaging. Their broad excitation profiles, superior photostability, and versatile surface chemistry functionalization capability further emphasize their suitability for such applications.[15-17] Based on spectral multiplexing method, QDs have found wide applications including monitoring the progression of cancer,[18, 19] identifying the cell types,[20] and assessing the genetic expression.[21] To realize the potential of QDs in temporally multiplexing, technical challenges must be addressed in preparing QDs with resolvable fluorescence lifetimes and selecting data acquisition and analysis methods suitable for lifetime scale of QDs.
Up to now, only a handful of studies have explored the feasibility of temporal multiplexing of QDs in solutions.[22, 23] However, the resolvability of their fluorescence lifetimes in solutions does not guarantee the resolvability of these QDs in cells and biological fluids, considering the sensitivity of their fluorescence lifetimes to pH and the biochemical species in local cellular environments.[24, 25] Until very recently, Cai's group successfully applied the type-II CdTe/CdS QDs with well separated fluorescence lifetimes (30-160 ns) for temporal multiplexing in cell imaging.[26] These type-II core/shell QDs exhibited prolonged fluorescence lifetimes due to the slow recombination of electrons and holes spatially separated in shell and core. However, this approach is difficult to be extended to other QDs due to the difficulty and challenge in preparing different types of highly luminescent type-II QDs.[27] Another possible approach to obtain QDs with widely separated fluorescence lifetimes is to combine QDs exhibiting band-edge PL with QDs exhibiting defect- or dopant-related PL. The slow recombination processes via intra-gap levels such as defects and dopants result in prolonging of the fluorescence lifetimes. For example, the CuInS2 and AgInS2 QDs have fluorescence lifetimes typically ranging from a hundred to a few hundred nanoseconds,[28-30] while the Cu-doped ZnS and Mn-doped ZnSe QDs show lifetimes up to microsecond and even millisecond scale.[13] These distinct temporal characteristics can be utilized to differentiate these QDs from the most commonly used Cd-based QDs with typical fluorescence lifetimes of several tens of nanoseconds.[31, 32] Herein, we attempted to examine the potential of this approach for temporally multiplexed imaging of cells. The CdSe/ZnS (CdSe) QDs and Zn-Ag-In-S (ZAIS) QDs are selected as models due to their widely-separated fluorescence lifetimes. Besides, these two QDs have wide tunability in emission wavelengths by varying size and compositions, which make them promising for simultaneous spectrally and temporally multiplexed imaging. By applying 4-plex QDs (green CdSe, red CdSe, green ZAIS, and red ZAIS QDs) with multi-lifetime and multi-emission in cell imaging, we obtained spectrally resolvable signals from green and red emitting QDs and temporally resolvable signals from CdSe and ZAIS QDs. For cells co-stained with different QDs and dye, the FLIM images showed spatially separated false-color patterns and can be split into channel images by using the software-based time gates. For the lifetime scale of QDs, the effect of data acquisition and data analysis methods on imaging results were discussed. Due to the rich availability of QDs with differing fluorescence lifetimes and emissions, the strategy used in this study can be easily extended to incorporate more QDs to maximize the number of measurable species in multiplexed imaging and sensing.
Materials and Methods
Synthesis and Surface Modification of Quantum Dots (QDs)
Organically-capped green-emitting and red-emitting CdSe/ZnS (ORG CdSe) QDs were obtained from mesolight and NN-Labs, respectively. The ORG CdSe QDs were transferred into the aqueous phase by a ligand exchange method described in our previous report.[33] Briefly, 1 mg of ORG CdSe QDs were washed with ethanol and mixed with 30 mg of mercaptosuccinic acid (MSA) in 5 ml of chloroform. After stirring for 5 min, 100 µl of 30% NH4OH solution and 900 µl of DI water were added into the reaction mixture, and the biphasic system solution was stirred for another 5 h. Afterwards, the QDs in aqueous phase were washed by centrifugation with ethanol and dispersed in 1 ml of DI water.
The synthesis of organically-capped Zn-Ag-In-S (ORG ZAIS) QDs was modified from previous protocols.[34, 35] Typically, 0.25 mmol of silver nitrate (AgNO3), 1 mmol of indium(III) acetate (In(OAc)3) and 3 mmol of stearic acid were mixed with 10 ml of 1-octadecene (ODE) and 2 ml of oleic acid in a flask. The reaction mixture was heated to 100 ˚C within 30 mins under stirring and nitrogen flow. After cooled down to 90 ˚C, 2 ml of dodecanethiol (DT) was added and the mixture was stirred for 3 min before the rapid injection of 1.5 mmol of sulfur (S) pre-heated in 3 mL of oleylamine. The Ag-In-S (AIS) QDs were allowed to grow for 5 min. Afterwards, 0.5 mmol of zinc stearate (ZnSt2) and S dispersed in ODE were injected into the reaction mixture and the temperature was raised to 150 ˚C. After 10 min, the ZAIS QDs were precipitated by centrifugation with ethanol and stored in chloroform. For preparing green-emitting ZAIS QDs, the Ag, In, Zn precursors were adjusted to 0.1 mmol, 0.4 mmol and 0.9 mmol. The Zn and S precursors were injected at 1 h interval for three times and the temperatures of the reaction mixture were respectively raised to 150 ˚C, 180 ˚C, and 220 ˚C after the first, second and third injection. For surface modification of ZAIS QDs, 1 mg of ORG ZAIS QDs dispersed in 100 µl of chloroform were injected into 1 ml aqueous solution of 46.5 mg cetyltrimethylammonium bromide (CTAB). After stirring at 70 °C to evaporate the chloroform, the resulting aqueous solution was added to the mixture of 4.5 ml of DI water and 12 µl of 0.2 M NaOH. Subsequently, 125 μl of tetraethyl orthosilicate (TEOS) was injected into the solution mixture, and the mixture was heated at 70 °C for 10 h under stirring. Afterwards, the silica modified ZAIS QDs were collected by centrifugation at 9000 rpm for 10 min. The QDs were washed with ethanol and water for five times to remove the impurities and dissolved in DI water with the assistance of sonication.
Characterization of QDs
Transmission electron microscopy (TEM) images were taken with a FEI TecnaiTM microscope at an accelerating voltage of 200 kV. Energy dispersive X-ray (EDX) spectra were obtained using a JSM-5600LV scanning electron microscope equipped with an EDX analyzer. UV-vis absorption spectra of the QDs were measured using a UV-2450 spectrometer (Shimadzu, Japan). Photoluminescence (PL) spectra of the QDs were recorded by a Fluorolog-3 Fluorometer (HORIBA Jobin Yvon, USA). The time-resolved PL decays of water-dispersible CdSe and ZAIS QDs were obtained by a DCS-120 Confocal Scanning Fluorescence Lifetime Imaging System (Becker & Hickl, Germany) operated at a single curve mode.
Incubation of QDs with Cells
RAW264.7 cells were cultured in Dulbecco's modified Eagles medium (DMEM) supplemented with 100 μg mL-1 penicillin, 100 μg mL-1 streptomycin and 10% fetal bovine serum (FBS) at 37 °C with 5% CO2. Before treating cells with QDs, cells were seeded onto the glass-bottom microwell dishes. The water-dispersible CdSe and ZAIS QDs were added into the culture media and the cells were incubated for another 4 h. After 4 h, the treated cells were washed with PBS for three times and fixed with 4% formaldehyde. 4′,6-Diamidino-2-phenylindole dihydrochloride (DAPI, Sigma) was used for nuclei staining. For cross-cell experiment, RAW264.7 cells separately treated with CdSe and ZAIS QDs in two dishes were washed, trypsinized and seeded onto one dish. After incubation for another 4 h, the cells were fixed with 4% formaldehyde.
Cell imaging studies
For spectral microscopy, the excitation intensity, exposure time for taking microscope images, and look-up tables (LUTs) of the acquired image were identical for [imaging the cells treated with green- and red-emitting QDs. For fluorescence lifetime imaging microscopy (FLIM), the cell images with 256 x 256 pixels and 256 time channels were taken at a film mode with 100x oil lens. 405 nm long pass filter was applied for acquiring fluorescent images. A 375 nm picosecond laser with 20 MHz repetition rate was used as the excitation source. HPM-100-40 high-speed hybrid detector was used to record the fluorescence signals by the control of DCC-100 detector controller. The fluorescence lifetimes in the FLIM image were calculated by fitting the decay curves using an incomplete decay model with the correction method (Supporting Information).
Results and Discussion
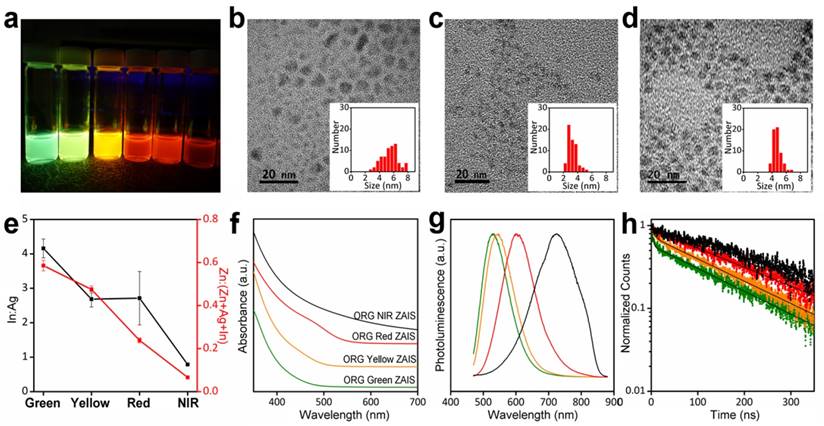
To achieve temporally and spectrally resolved fluorescence signals, the CdSe/ZnS QDs (shortly as CdSe QDs) and Zn-Ag-In-S (ZAIS) QDs with wide PL tunability and distinct fluorescence lifetimes were selected as QDs models for multiplexed cell imaging. Organically-capped (ORG) CdSe QDs with different emission wavelengths are commercially available and ORG ZAIS QDs with tunable PL were prepared by using the composition control strategy.[29] By varying the Ag:In feed ratio in nucleation stage and alloying Zn and S at elevated temperature, we obtained ORG ZAIS QDs with green to near-infrared (NIR) emissions under UV excitation (Figure 1a). The quantum yields (QYs) are as high as 33.8-66.7% for red- and green-emitting ZAIS QDs and 11.2% for NIR-emitting ZAIS QDs. The size of green-, orange- and NIR-emitting QDs estimated from transmission electron microscopy (TEM) images are 5.5±1.2, 3.2±0.6, and 4.7±0.6 nm, respectively (Figure 1b, c, d). The size broadening observed in green-emitting ZAIS QDs may result from the non-homogeneous alloying of Zn or Ostwald ripening during the prolonged reaction time. Energy dispersive X-ray (EDX) results confirmed the compositional variation in the QDs synthesized with different precursor ratios (Figure 1e). The TEM and EDX results combinedly indicate that the wavelength differences of ZAIS QDs mainly result from the compositional variation rather than the size changes. By decreasing the Ag:In ratio or increasing the Zn content, the absorption and emission spectra of the ZAIS QDs showed a pronounced blue shift (Figure 1f, g), which was attributed to the bandgap widening effect upon composition control.[30, 36] At the same time, a shortening of the fluorescence lifetime from 243 to 155 ns (average fluorescence lifetime calculated by bi-exponential fitting) was observed in ZAIS QDs (Figure 1h). Despite this variation, all of these fluorescence lifetimes at the scale of hundreds of nanoseconds are substantially different from those of the typically used CdSe QDs, suggesting the potential of combining these ZAIS QDs with CdSe QDs for temporally multiplexed imaging.
Characterization of organically-capped Zn-Ag-In-S (ORG ZAIS) QDs with different composition ratios. (a) Digital photograph of ZAIS QDs with different Ag:In ratio and Zn content under UV light. TEM images and size distributions of (b) green-emitting, (c) red-emitting and (d) NIR-emitting ZAIS QDs. (e) Composition ratio of ZAIS QDs determined by EDX. (f) UV-vis absorption and (g) PL spectra of green (olive line), yellow (orange line), red (red line) and NIR (black line) emitting ZAIS QDs with different composition ratio. (h) Time-resolved PL decays (dots) and bi-exponential fitting curves (black line) of the same ZAIS QDs.
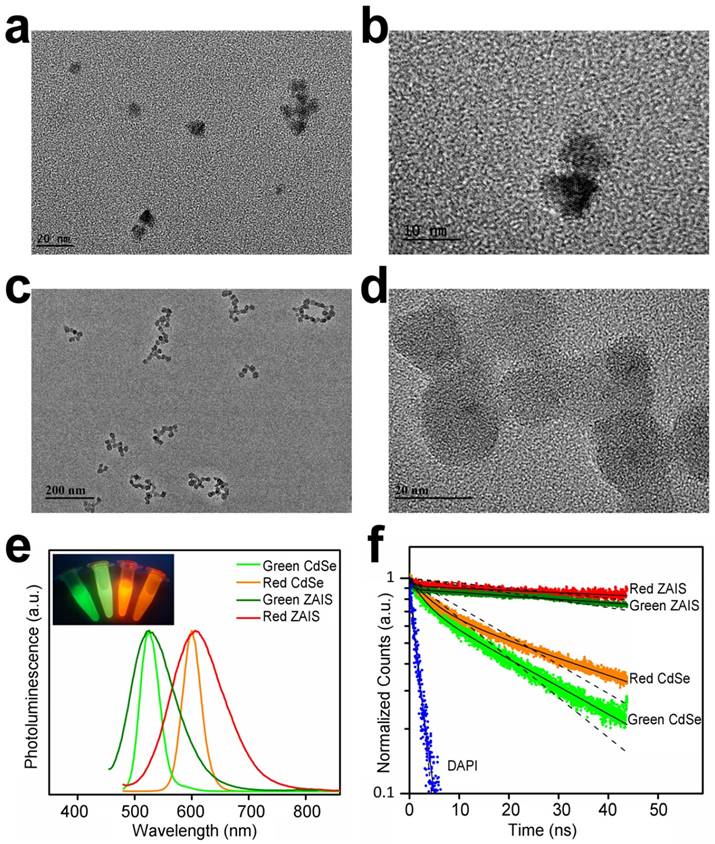
The green- and red-emitting CdSe and ZAIS QDs were selected as 2-dimensional encoded labels with different optical characteristics in spectral and time domain. For biomedical imaging studies, organic-aqueous phase transfer of these QDs is required to make them water dispersible. The ORG CdSe QDs were transferred into the aqueous phase by using the mercaptosuccinic acid (MSA) as surface ligands. The obtained MSA-capped CdSe QDs remained the morphology and crystallinity of the organic-soluble CdSe QDs (Figure 2a, 2b, S1). The ORG ZAIS QDs were rendered hydrophillic by encapsulating them with a silica shell. TEM images show that the ZAIS QDs were embedded within the silica shell of ~10 nm thickness (Figure 2c, 2d). The QYs of the red- and green-emitting ZAIS QDs are 11.5%-21.4% after transferring them to aqueous phase. The spectral and temporal optical properties of the surface modified CdSe and ZAIS QDs in solutions were compared through analysis of their PL spectra and time-resolved PL decays. From Figure 2e, the PL spectra could be utilized to differentiate the green emitting with red emitting QDs. For green CdSe and green ZAIS QDs (or red CdSe and red ZAIS QDs), their absorption (Figure S2) and PL spectra are severely overlapped, making it difficult to distinguish their emissions by optical filters or visible eyes (Figure 2e, inset).
Characterization of water-dispersible CdSe and ZAIS QDs. TEM images of (a, b) MSA-capped CdSe QDs and (c, d) silica-coated ZAIS QDs. (e) PL spectra of 4-plex CdSe and ZAIS QDs dispersed in water. Inset is a digital photograph showing the aqueous dispersions of these QDs under UV light (left to right: green CdSe QDs, green ZAIS QDs, red CdSe QDs, red ZAIS QDs). (f) Time-resolved PL decays of DAPI in cell nuclei, green and red emitting CdSe and ZAIS QDs dispersed in water. The PL decays were fitted to mono-exponential (dash line) and bi-exponential (solid line) curves.
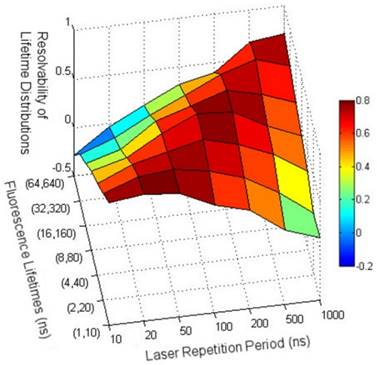
Despite the spectral similarity between these QDs, they demonstrate a significant difference in their time-resolved PL decay profiles (Figure 2f). Moreover, both of their PL decay profiles show a much lower decay rate compared to that of 4,6-diamidino-2-phenylindole dihydrochloride (DAPI), a dye model used in this study. It should be noted that, due to the high frequency (20 MHz) of the applied laser, the recorded decays are incomplete fluorescence decays, containing the fluorescence decay upon excitation of current pulse and the decay tails upon excitation of previous pulses. This incomplete effect caused by the high excitation frequency could skew the decay curves and blur the difference between the fast and slow decays. A lower frequency excitation will allow the acquisition of more complete fluorescence decays, but limit the number of photons that can be collected in a given time. The smaller photon number means a smaller signal-to-noise ratio (SNR) and results in broad distributions of the calculated lifetimes, thereby increasing the difficulty of resolving the different fluorescence lifetimes. The effect of excitation frequency on the resolvability of the fluorescence lifetime distributions was investigated by simulation (Figure 3). For two specific lifetimes, lengthening of the laser repetition period (i.e., decrease of the laser frequency) first increases the resolvability due to the narrowing of the long lifetime distributions while further lengthening of the laser repetition periods leads to a lower resolvability due to the broadening of both lifetime distributions (Figure 3 and Figure S3).
Simulation results on the effects of the laser repetition rates (x axis) and fluorescence lifetime scale (y axis) on the resolvability of the fluorescence lifetime distributions (z axis). The first and second number of the y tick labels is respectively short and long fluorescence lifetime values for generating decay curves. The resolvability of the two lifetime distributions was calculated as the distance between the right tail of the short lifetime distribution and the left tail of the long lifetime distribution relative to the distance between the mean of two lifetimes.
As expected, the optimal laser repetition periods are larger for resolving longer fluorescence lifetimes. In this study, the applied 20 MHz laser has a repetition period of 50 ns, which increases the resolvability of the PL decays of DAPI and CdSe QDs and retains the difference between the PL decays of the CdSe and ZAIS QDs. To incorporate other QDs with longer fluorescence lifetimes (e.g. Mn- or Cu- doped QDs) in this system, a longer excitation repetition rate was suggested to be applied, either by using a lower frequency laser or by modulating the on-time of laser.[37] After curve fitting of the collected decays, an incomplete decay model with corrections was used to estimate the fluorescence lifetimes (Supporting Information). The fluorescence lifetimes calculated from the mono-exponential fitting curves are 1.93, 23.5, 32.4, 127, and 188 ns respectively for DAPI in cell nuclei, green CdSe, red CdSe, green ZAIS, and red ZAIS QDs in water. Although the mono-exponential fitting is not a tight fitting model for these multi-exponential fluorescence decays in comparison with the bi-exponential fitting (Figure S4), it is recommended to be used in cell imaging because it is more robust when there are limited numbers of photons for curve fitting.[38] Additionally, the corrected bi-exponential fitting model proved that the lifetime values obtained by the mono-exponential fitting model are reasonable (Table S2).
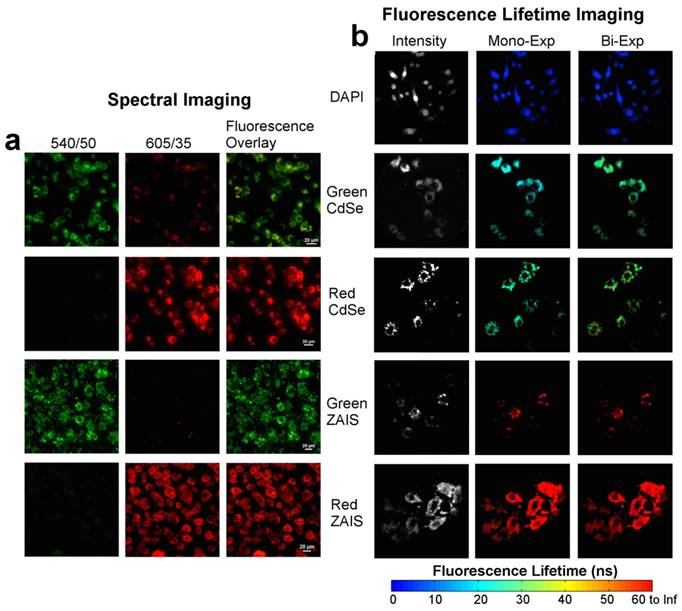
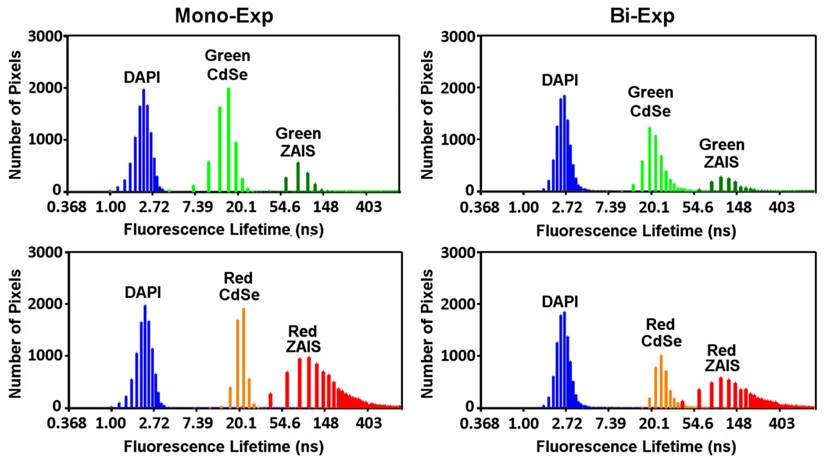
To examine the potential of the 2-dimensional encoded QDs for spectrally and temporally multiplexed imaging, the cells treated separately with these QDs were imaged by spectral microscopy and fluorescence lifetime imaging microscopy (FLIM). In spectral imaging, the fluorescent signals from the green and red emitting QDs were resolved by using spectrally matched filters (Figure 4a). In FLIM imaging, the fluorescence decays in each pixel were fitted to mono-exponential curves and the calculated lifetimes were correlated with the hue component of a hue-saturation-value (HSV) image. The resulting false-color images show clear distinction between signals from CdSe QDs and ZAIS QDs (Figure 4b, middle column). The fluorescence lifetimes of these QDs in cells could be further discriminated from those of DAPI, which shows the potential of combining these QDs with organic dyes to achieve three levels of fluorescence lifetimes in temporally multiplexed imaging. The broad distribution of the fluorescence lifetimes (Figure 5, left panel) is considered to be a combined result of the lifetime variation of dye and QDs in cells,[24, 25, 39] noise in data collection, and error in data analysis. Carlini and Nadeau applied CdSe QDs for FLIM imaging and found their fluorescence lifetimes varied in different cellular regions, which are affected by the pH level and redox conditions of local environment.[24] Similarly, Orte et al. have observed that the fluorescence lifetimes of CdSe QDs varied from 8.7 to 15.4 ns when the solution pH was increased from 5 to 9, and they even utilized this property for sensing pH in live cells.[25] However, despite the lifetime variation, the emission of DAPI, CdSe and ZAIS QDs can be well distinguished by their fluorescence lifetimes when applying an appropriate lifetime scale (0-60 ns in Figure 4). The subtle variation of fluorescence lifetimes in different cellular regions can be observed if a smaller and more specific lifetime scale is applied to display the image. As an example, the same image of cells treated with red CdSe QDs were displayed in a lifetime scale of 17-25 ns and showed observable variation in fluorescence lifetimes from different regions (Figure S5). By utilizing the sensitivity of fluorescence lifetimes, the variation could be used for probing cellular environment. This indicated the potential of using FLIM imaging of QDs for simultaneously temporally multiplexed imaging and sensing, which more extensively exploits the available information. Upon comparing to mono-exponential fitting, the bi- exponential fitting generated less resolvable FLIM images of the applied dye and QDs (right columns of Figure 4 and Figure 5). This may result from a higher instability of bi-exponential fitting under the condition of limited photon numbers [38] and the over-weighting of long lifetime component when calculating the intensity averaged lifetime.[24, 40] Therefore, for our FLIM image with limited photons numbers (<104 photons in each pixel), mono-exponential model is recommended for curve fitting and lifetime calculations. Overall, these cell imaging results demonstrate the potential of using the 2-dimensional encoded QDs for spectrally and temporally multiplexed imaging.
(a) Spectral imaging and (b) fluorescence lifetime imaging of RAW264.7 cells separately treated with DAPI, green CdSe, red CdSe, green ZAIS, and red ZAIS QDs. The fluoerscence lifetimes were obtained by mono- or bi- exponential fitting and mapped to a false-color scale from 0 ns (blue) to 60 ns (red).
Fluorescence lifetime distributions of DAPI, green CdSe, green ZAIS, red CdSe, and red ZAIS QDs in FLIM images obtained by mono-exponential (left panel) and bi-exponential (right panel) fitting.
In spectrally multiplexed imaging, the channel splitting is achieved by using a series of spectrally matched optical filters. In temporally multiplexed imaging, the channel splitting can be carried out by using software or programmable scripts, thus can be easily customized according to the specific temporal characteristics of the fluorescent probes. Herein, by deliberately introducing any two of DAPI, red CdSe and red ZAIS QDs in a same FLIM image, we obtained three mock scenarios for temporally multiplexed imaging. The obtained fluorescence lifetime distributions (Figure 5) were used to determine the time gates for channel splitting of these temporally multiplexed images. Accordingly, 10 ns was set as the time gate to separate the signals from DAPI and red CdSe QDs and 40 ns was selected to separate signals from red CdSe and ZAIS QDs. By using the software-based time gates, the FLIM images containing a mixture of fluorescence signals were split into three channel images (Figure 6). In original FLIM images, the cells co-stained with DAPI and either type of QDs exhibit a false-color pattern of round nuclei encircled by peripheral cell regions. The cell nuclei were displayed in 0-10 ns channel image, while the peripheral regions were mostly present in 10-40 ns channel image for cells treated with red CdSe QDs and in >40 ns channel image for cells treated with red ZAIS QDs.
Temporally multiplexed imaging of the RAW264.7 cells co-stained with DAPI and red CdSe QDs (top row), DAPI and red ZAIS QDs (middle row), and the cells separately treated with red CdSe and red ZAIS QDs, seeded and cultured in a same dish (bottom row). The original FLIM images (left column) were split into 0-10 ns, 10-40 ns, and > 40 ns channel images as indicated.
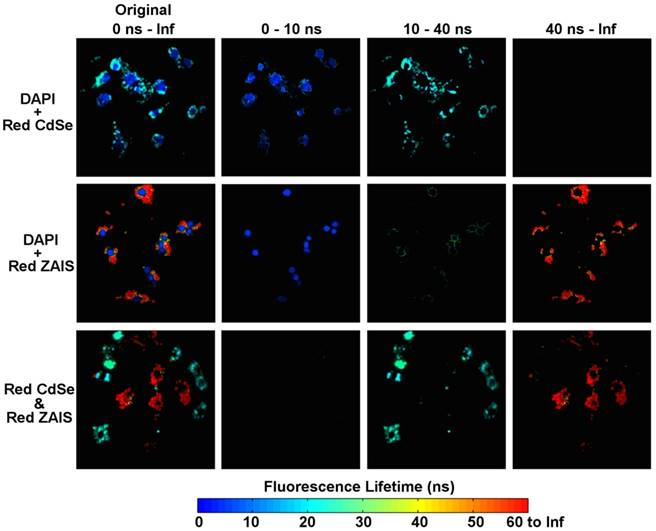
This result indicates that in multiplexed cell imaging, the emissions from QDs and dyes with distinct lifetimes can be temporally resolved. Also, we noticed that a perinuclear, ring-like pattern showing lifetimes in between those of the DAPI and QDs, which was displayed in 0-10 ns channel image for cells treated with red CdSe QDs and DAPI, and in 10-40 ns image channel for cells treated with ZAIS QDs and DAPI. According to results in Figure 4, DAPI only or QDs only cannot generate such a pattern with fluorescence lifetimes in the mediate range. Therefore, this pattern is most likely to arise from the co-existence of DAPI and QD signals with comparative intensity in the same pixel. To examine the resolvability of two QDs in a same cell image, we performed a cross-cell experiment [41] by co-incubating two group of cells, either pretreated with red CdSe or red ZAIS QDs, in a same dish for FLIM imaging. The resulting FLIM images show a spatially separated pattern of cells with different false colors, which is expected when no or very few QDs were transferred between neighboring cells. After channel splitting, the blue-green pixels in 10-40 ns channel image and red pixels in >40 ns channel image are respectively attributed to emissions from red CdSe and red ZAIS QDs. This result conceptually proved that the emissions of DAPI, CdSe and ZAIS QDs can be resolved from temporally multiplexed cell imaging. The same temporal multiplexing technique can improve the reliability of distinguishing the fluorescent probes with similar spectral wavelengths but distinct fluorescent lifetimes.
Conclusions
In summary, our proof-of-concept study demonstrates the feasibility of spectrally and temporally multiplexed imaging of cells by using 4-plex QDs with different emission wavelengths and widely differing fluorescent lifetimes. Green emitting CdSe and ZAIS QDs were differentiated from red emitting CdSe and ZAIS QDs by spectral imaging. CdSe and ZAIS QDs with similar spectral wavelengths were well resolved by FLIM imaging when taken up by cells, which were attributed to the substantial difference in their fluorescence lifetimes. Future work is needed to extend this technique to a variety of QDs showing magnitude difference of fluorescence lifetimes. The excitation frequency and data analysis method should be carefully designed according to the lifetime scale of the applied QDs. Combining a pool of spectrally and temporally distinct fluorescent probes, one can expect to increase or even multiply the number of measurable species in multiplexed imaging and sensing applications. Such technique will serve as a valuable tool for future nanomedicine and biophotonics research.
Supplementary Material
Supplementary figures and tables.
Acknowledgements
This study was supported by the Ministry of Education, Singapore (Tier 2 MOE2010-T2-2-010 (M4020020.040 ARC2/11)), NTU-NHG Innovation Collaboration Grant (No. M4061202.040), A*STAR Science and Engineering Research Council (No. M4070176.040), NTU-A*STAR Silicon Technologies, Centre of Excellence under the program grant No. 11235100003, NEWRI Seed Grant, and School of Electrical and Electronic Engineering at Nanyang Technological University.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Choi HM, Chang JY, Trinh LA, Padilla JE, Fraser SE, Pierce NA. Programmable in situ amplification for multiplexed imaging of mRNA expression. Nat Biotechnol. 2010;28:1208-12
2. Kobayashi H, Longmire MR, Ogawa M, Choyke PL, Kawamoto S. Multiplexed imaging in cancer diagnosis: applications and future advances. Lancet Oncol. 2010;11:589-95
3. Ding H, Yong K-T, Law W-C, Roy I, Hu R, Wu F. et al. Non-invasive tumor detection in small animals using novel functional Pluronic nanomicelles conjugated with anti-mesothelin antibody. Nanoscale. 2011;3:1813-22
4. Kumar R, Ding H, Hu R, Yong K-T, Roy I, Bergey EJ. et al. In vitro and in vivo optical imaging using water-dispersible, noncytotoxic, luminescent, silica-coated quantum rods. Chem Mater. 2010;22:2261-7
5. Hoffman RM. Use of fluorescent proteins and color-coded imaging to visualize cancer cells with different genetic properties. Cancer Metastasis Rev. 2016;35:5-19
6. Yong K-T, Wang Y, Roy I, Rui H, Swihart MT, Law W-C. et al. Preparation of quantum dot/drug nanoparticle formulations for traceable targeted delivery and therapy. Theranostics. 2012;2:681-94
7. Hoffmann K, Behnke T, Drescher D, Kneipp J, Resch-Genger U. Near-infrared-emitting nanoparticles for lifetime-based multiplexed analysis and imaging of living cells. ACS Nano. 2013;7:6674-84
8. Hoffmann K, Behnke T, Grabolle M, Resch-Genger U. Nanoparticle-encapsulated vis-and NIR-emissive fluorophores with different fluorescence decay kinetics for lifetime multiplexing. Anal Bioanal Chem. 2014;406:3315-22
9. Mandal G, Darragh M, Andrew Wang Y, Heyes C-D. Cadmium-free quantum dots as time-gated bioimaging probes in highly-autofluorescent human breast cancer cells. Chem Commun. 2013;49:624-6
10. Gu L, Hall DJ, Qin Z, Anglin E, Joo J, Mooney DJ. et al. In vivo time-gated fluorescence imaging with biodegradable luminescent porous silicon nanoparticles. Nat Commun. 2013;4:2326
11. Vereb G, Jares-Erijman E, Selvin PR, Jovin TM. Temporally and spectrally resolved imaging microscopy of lanthanide chelates. Biophys J. 1998;74:2210-22
12. Lu Y, Lu J, Zhao J, Cusido J, Raymo FM, Yuan J. et al. On-the-fly decoding luminescence lifetimes in the microsecond region for lanthanide-encoded suspension arrays. Nat Commun. 2014;5:3741
13. Wu P, Yan X-P. Doped quantum dots for chemo/biosensing and bioimaging. Chem Soc Rev. 2013;42:5489-521
14. Yong K-T, Roy I, Law W-C, Hu R. Synthesis of cRGD-peptide conjugated near-infrared CdTe/ZnSe core-shell quantum dots for in vivo cancer targeting and imaging. Chem Commun. 2010;46:7136-8
15. Xu G, Zeng S, Zhang B, Swihart MT, Yong K-T, Prasad PN. New generation cadmium-free quantum dots for biophotonics and nanomedicine. Chem Rev. 2016;116:12234-327
16. Erogbogbo F, Chang C-W, May JL, Liu L, Kumar R, Law W-C. et al. Bioconjugation of luminescent silicon quantum dots to gadolinium ions for bioimaging applications. Nanoscale. 2012;4:5483-9
17. Hu S, Zeng S, Zhang B, Yang C, Song P, Danny TJH. et al. Preparation of biofunctionalized quantum dots using microfluidic chips for bioimaging. Analyst. 2014;139:4681-90
18. Liu J, Lau SK, Varma VA, Moffitt RA, Caldwell M, Liu T. et al. Molecular mapping of tumor heterogeneity on clinical tissue specimens with multiplexed quantum dots. ACS Nano. 2010;4:2755-65
19. Peng C-W, Liu X-L, Chen C, Liu X, Yang X-Q, Pang D-W. et al. Patterns of cancer invasion revealed by QDs-based quantitative multiplexed imaging of tumor microenvironment. Biomaterials. 2011;32:2907-17
20. He GS, Zheng Q, Yong K-T, Erogbogbo F, Swihart MT, Prasad PN. Two-and three-photon absorption and frequency upconverted emission of silicon quantum dots. Nano Lett. 2008;8:2688-92
21. Kang WJ, Chae JR, Cho YL, Lee JD, Kim S. Multiplex imaging of single tumor cells using quantum-dot-conjugated aptamers. Small. 2009;5:2519-22
22. Chen C, Zhang P, Gao G, Gao D, Yang Y, Liu H. et al. Near-infrared-emitting two-dimensional codes based on lattice-strained core/(doped) shell quantum dots with long fluorescence lifetime. Adv Mater. 2014;26:6313-7
23. Grabolle M, Kapusta P, Nann T, Shu X, Ziegler J, Resch-Genger U. Fluorescence lifetime multiplexing with nanocrystals and organic labels. Anal Chem. 2009;81:7807-13
24. Carlini L, Nadeau JL. Uptake and processing of semiconductor quantum dots in living cells studied by fluorescence lifetime imaging microscopy (FLIM). Chem Commun. 2013;49:1714-6
25. Orte A, Alvarez-Pez JM, Ruedas-Rama MJ. Fluorescence lifetime imaging microscopy for the detection of intracellular pH with quantum dot nanosensors. ACS Nano. 2013;7:6387-95
26. Zhang L, Chen C, Li W, Gao G, Gong P, Cai L. Living cell multilifetime encoding based on lifetime-tunable lattice-strained quantum dots. ACS Appl Mater Interfaces. 2016;8:13187-91
27. Donegá C-M. Synthesis and properties of colloidal heteronanocrystals. Chem Soc Rev. 2011;40:1512-46
28. Kim Y-K, Ahn S-H, Chung K, Cho Y-S, Choi C-J. The photoluminescence of CuInS2 nanocrystals: effect of non-stoichiometry and surface modification. J Mater Chem. 2012;22:1516-20
29. Zhang B, Wang Y, Yang C, Hu S, Gao Y, Zhang Y. et al. The composition effect on the optical properties of aqueous synthesized Cu-In-S and Zn-Cu-In-S quantum dot nanocrystals. Phys Chem Chem Phys. 2015;17:25133-41
30. Xiang W, Xie C, Wang J, Zhong J, Liang X, Yang H. et al. Studies on highly luminescent AgInS2 and Ag-Zn-In-S quantum dots. J Alloys Compd. 2014;588:114-21
31. Resch-Genger U, Grabolle M, Cavaliere-Jaricot S, Nitschke R, Nann T. Quantum dots versus organic dyes as fluorescent labels. Nat Methods. 2008;5:763-75
32. Issac A, Jin S, Lian T. Intermittent electron transfer activity from single CdSe/ZnS quantum dots. J Am Chem Soc. 2008;130:11280-1
33. Zhang B, Hu R, Wang Y, Yang C, Liu X, Yong K-T. Revisiting the principles of preparing aqueous quantum dots for biological applications: the effects of surface ligands on the physicochemical properties of quantum dots. RSC Adv. 2014;4:13805-16
34. Yoon HC, Oh JH, Ko M, Yoo H, Do YR. Synthesis and characterization of green Zn-Ag-In-S and red Zn-Cu-In-S quantum dots for ultrahigh color quality of down-converted white LEDs. ACS Appl Mater Interfaces. 2015;7:7342-50
35. Liu L, Hu R, Law W-C, Roy I, Zhu J, Ye L. et al. Optimizing the synthesis of red-and near-infrared CuInS2 and AgInS2 semiconductor nanocrystals for bioimaging. Analyst. 2013;138:6144-53
36. Hamanaka Y, Yukitoki D, Kuzuya T. Structural transformation and photoluminescence modification of AgInS2 nanoparticles induced by ZnS shell formation. Appl Phys Express. 2015;8:095001
37. Becker W, Su B, Bergmann A, Weisshart K, Holub O. Simultaneous fluorescence and phosphorescence lifetime imaging. SPIE BiOS: International Society for Optics and Photonics. 2011;7903:790320-7
38. Spriet C, Trinel D, Riquet F, Vandenbunder B, Usson Y, Heliot L. Enhanced FRET contrast in lifetime imaging. Cytometry A. 2008;73:745-53
39. Sanchez WY, Song Z, Becker W, Koenig K, Roberts MS. Fluorescence lifetime imaging for diagnostic and therapeutic intravital microscopy. In: (ed.) Weigert R. Advances in Intravital Microscopy: From Basic to Clinical Research. Dordrecht: Springer Netherlands. 2014:371-418
40. Berberan-Santos MN, Bodunov EN, Valeur B. Mathematical functions for the analysis of luminescence decays with underlying distributions 1. Kohlrausch decay function (stretched exponential). Chem Phys. 2005;315:171-82
41. Slowing II, Vivero-Escoto JL, Zhao Y, Kandel K, Peeraphatdit C, Trewyn BG. et al. Exocytosis of mesoporous silica nanoparticles from mammalian cells: from asymmetric cell-to-cell transfer to protein harvesting. Small. 2011;7:1526-32
Author contact
Corresponding authors: Prof. Liwei Liu, E-mail: liulwedu.cn; Prof. Junle Qu, E-mail: jlquedu.cn; Prof. Ken-Tye Yong, E-mail: ktyongedu.sg

 Global reach, higher impact
Global reach, higher impact